Analysis of long non-coding RNA expression profiles in clear cell renal cell carcinoma
- PMID: 28928816
- PMCID: PMC5588171
- DOI: 10.3892/ol.2017.6563
Analysis of long non-coding RNA expression profiles in clear cell renal cell carcinoma
Abstract
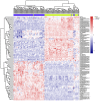
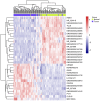
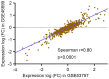
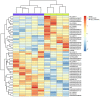
To investigate the expression patterns of long non-coding RNAs (lncRNAs) in clear cell renal cell carcinoma (ccRCC) and in metastatic renal cell carcinoma (RCC), the present study downloaded three human exon arrays available from the public Gene Expression Omnibus. The probes of the human exon arrays were re-annotated and the probes uniquely mapping to lncRNAs were retained at the gene level. Following the analysis of GSE53757 and GSE46699, which contained paired ccRCC cancer and normal adjacent tissue samples, 32 differentially expressed lncRNAs (adjusted P<0.01) in ccRCC were identified. Various lncRNAs, including ENSG00000177133, NR_024418, T-cell leukemia/lymphoma 6 (TCL6), growth arrest-specific transcript 5, deleted in lymphocytic leukemia 2, colorectal neoplasia differentially expressed (CRNDE) and MIR155HG, have been reported to be abnormally expressed in cancers. Of these genes, NR_24418 and TCL6 have been reported to be associated with ccRCC. Following analysis of GSE47352, which contained 4 primary metastatic and 5 non-metastatic tumor samples, the 50 top differentially expressed lncRNAs were identified in metastatic ccRCC (Mann-Whitney U test, P<0.05). Comparison with the ccRCC associated lncRNAs revealed that the lncRNA CRNDE demonstrated an increased expression in ccRCC and metastatic ccRCC samples, which suggested that CRNDE is important in the progression of ccRCC. The lncRNA ENSG00000244020 was decreased in ccRCC and metastatic ccRCC, suggesting that silencing of ENSG00000244020 may be important in ccRCC development. Overall, a set of lncRNAs was identified as differentially expressed in ccRCC and metastatic ccRCC, providing potential candidates for the discovery of novel cancer biomarkers and therapeutic targets to improve diagnosis and therapy in RCC.
Keywords: clear cell renal cell carcinoma; data mining; long non-coding RNA; microarray analysis.
Figures
Similar articles
-
Decreased TCL6 expression is associated with poor prognosis in patients with clear cell renal cell carcinoma.Oncotarget. 2017 Jan 24;8(4):5789-5799. doi: 10.18632/oncotarget.11011. Oncotarget. 2017. PMID: 27494890 Free PMC article.
-
Dysregulation of Long Non-coding RNAs and mRNAs in Plasma of Clear Cell Renal Cell Carcinoma Patients Using Microarray and Bioinformatic Analysis.Front Oncol. 2020 Nov 27;10:559730. doi: 10.3389/fonc.2020.559730. eCollection 2020. Front Oncol. 2020. PMID: 33330027 Free PMC article.
-
Analysis of long non-coding RNA expression profiles in gastric cancer.World J Gastroenterol. 2013 Jun 21;19(23):3658-64. doi: 10.3748/wjg.v19.i23.3658. World J Gastroenterol. 2013. PMID: 23801869 Free PMC article.
-
Current Concepts of Non-Coding RNAs in the Pathogenesis of Non-Clear Cell Renal Cell Carcinoma.Cancers (Basel). 2019 Oct 17;11(10):1580. doi: 10.3390/cancers11101580. Cancers (Basel). 2019. PMID: 31627266 Free PMC article. Review.
-
A review on the role of long non-coding RNA and microRNA network in clear cell renal cell carcinoma and its tumor microenvironment.Cancer Cell Int. 2023 Feb 2;23(1):16. doi: 10.1186/s12935-023-02861-6. Cancer Cell Int. 2023. PMID: 36732762 Free PMC article. Review.
Cited by
-
MIR155HG is a prognostic biomarker and associated with immune infiltration and immune checkpoint molecules expression in multiple cancers.Cancer Med. 2019 Dec;8(17):7161-7173. doi: 10.1002/cam4.2583. Epub 2019 Sep 30. Cancer Med. 2019. PMID: 31568700 Free PMC article.
-
Colorectal neoplasia differentially expressed: a long noncoding RNA with an imperative role in cancer.Onco Targets Ther. 2018 Jun 29;11:3755-3763. doi: 10.2147/OTT.S162754. eCollection 2018. Onco Targets Ther. 2018. PMID: 29988699 Free PMC article. Review.
-
The Role of Long Noncoding RNA (lncRNAs) Biomarkers in Renal Cell Carcinoma.Int J Mol Sci. 2022 Dec 30;24(1):643. doi: 10.3390/ijms24010643. Int J Mol Sci. 2022. PMID: 36614082 Free PMC article. Review.
-
CRNDE: an oncogenic long non-coding RNA in cancers.Cancer Cell Int. 2020 May 12;20:162. doi: 10.1186/s12935-020-01246-3. eCollection 2020. Cancer Cell Int. 2020. PMID: 32435153 Free PMC article. Review.
-
[Comprehensive analysis of the aberrantly expressed profiles of lncRNAs, miRNAs and the regulation network of the associated ceRNAs in clear cell renal cell carcinoma].Sheng Wu Yi Xue Gong Cheng Xue Za Zhi. 2019 Apr 25;36(2):267-273. doi: 10.7507/1001-5515.201801057. Sheng Wu Yi Xue Gong Cheng Xue Za Zhi. 2019. PMID: 31016944 Free PMC article. Chinese.
References
LinkOut - more resources
Full Text Sources
Other Literature Sources
