Detection of Bacterial Pathogens from Broncho-Alveolar Lavage by Next-Generation Sequencing
- PMID: 28930150
- PMCID: PMC5618659
- DOI: 10.3390/ijms18092011
Detection of Bacterial Pathogens from Broncho-Alveolar Lavage by Next-Generation Sequencing
Abstract
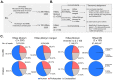
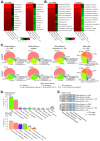
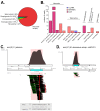
The applications of whole-metagenome shotgun sequencing (WMGS) in routine clinical analysis are still limited. A combination of a DNA extraction procedure, sequencing, and bioinformatics tools is essential for the removal of human DNA and for improving bacterial species identification in a timely manner. We tackled these issues with a broncho-alveolar lavage (BAL) sample from an immunocompromised patient who had developed severe chronic pneumonia. We extracted DNA from the BAL sample with protocols based either on sequential lysis of human and bacterial cells or on the mechanical disruption of all cells. Metagenomic libraries were sequenced on Illumina HiSeq platforms. Microbial community composition was determined by k-mer analysis or by mapping to taxonomic markers. Results were compared to those obtained by conventional clinical culture and molecular methods. Compared to mechanical cell disruption, a sequential lysis protocol resulted in a significantly increased proportion of bacterial DNA over human DNA and higher sequence coverage of Mycobacterium abscessus, Corynebacterium jeikeium and Rothia dentocariosa, the bacteria reported by clinical microbiology tests. In addition, we identified anaerobic bacteria not searched for by the clinical laboratory. Our results further support the implementation of WMGS in clinical routine diagnosis for bacterial identification.
Keywords: broncho-alveolar lavage; clinical metagenomics; whole-metagenome shotgun.
Conflict of interest statement
The authors declare that they have no competing interests.
Figures
References
-
- Hasman H., Saputra D., Sicheritz-Ponten T., Lund O., Svendsen C.A., Frimodt-Møller N., Aarestrup F.M. Rapid Whole-genome sequencing for detection and characterization of microorganisms directly from clinical samples. J. Clin. Microbiol. 2014;52:139–146. doi: 10.1128/JCM.02452-13. - DOI - PMC - PubMed
-
- Schmidt K., Mwaigwisya S., Crossman L.C., Doumith M., Munroe D., Pires C., Khan A.M., Woodford N., Saunders N.J., Wain J., et al. Identification of bacterial pathogens and antimicrobial resistance directly from clinical urines by nanopore-based metagenomic sequencing. J. Antimicrob. Chemother. 2017;72:104–114. doi: 10.1093/jac/dkw397. - DOI - PubMed
-
- Naccache S.N., Peggs K.S., Mattes F.M., Phadke R., Garson J.A., Grant P., Samayoa E., Federman S., Miller S., Lunn M.P., et al. Diagnosis of neuroinvasive astrovirus infection in an immunocompromised adult with encephalitis by unbiased next-generation sequencing. Clin. Infect. Dis. 2015;60:919–923. doi: 10.1093/cid/ciu912. - DOI - PMC - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

