The non-canonical poly(A) polymerase FAM46C acts as an onco-suppressor in multiple myeloma
- PMID: 28931820
- PMCID: PMC5606997
- DOI: 10.1038/s41467-017-00578-5
The non-canonical poly(A) polymerase FAM46C acts as an onco-suppressor in multiple myeloma
Abstract
FAM46C is one of the most frequently mutated genes in multiple myeloma. Here, using a combination of in vitro and in vivo approaches, we demonstrate that FAM46C encodes an active non-canonical poly(A) polymerase which enhances mRNA stability and gene expression. Reintroduction of active FAM46C into multiple myeloma cell lines, but not its catalytically-inactive mutant, leads to broad polyadenylation and stabilization of mRNAs strongly enriched with those encoding endoplasmic reticulum-targeted proteins and induces cell death. Moreover, silencing of FAM46C in multiple myeloma cells expressing WT protein enhance cell proliferation. Finally, using a FAM46C-FLAG knock-in mouse strain, we show that the FAM46C protein is strongly induced during activation of primary splenocytes and that B lymphocytes isolated from newly generated FAM46C KO mice proliferate faster than those isolated from their WT littermates. Concluding, our data clearly indicate that FAM46C works as an onco-suppressor, with the specificity for B-lymphocyte lineage from which multiple myeloma originates. FAM46C is one of the most frequently mutated genes in multiple myeloma (MM), but its molecular function remains unknown. Here the authors show that FAM46C is a poly(A) polymerase and that loss of function of FAM46C drives multiple myeloma through the destabilisation of ER response transcripts.
Conflict of interest statement
The authors declare no competing financial interests.
Figures




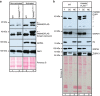



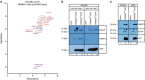
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials

