Hox gene cluster of the ascidian, Halocynthia roretzi, reveals multiple ancient steps of cluster disintegration during ascidian evolution
- PMID: 28932414
- PMCID: PMC5602962
- DOI: 10.1186/s40851-017-0078-3
Hox gene cluster of the ascidian, Halocynthia roretzi, reveals multiple ancient steps of cluster disintegration during ascidian evolution
Abstract
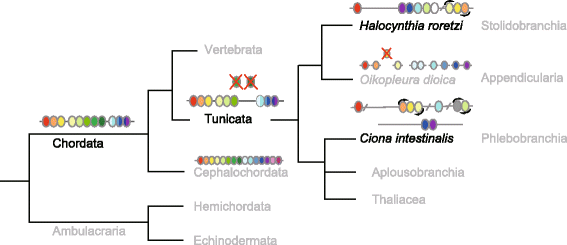
Background: Hox gene clusters with at least 13 paralog group (PG) members are common in vertebrate genomes and in that of amphioxus. Ascidians, which belong to the subphylum Tunicata (Urochordata), are phylogenetically positioned between vertebrates and amphioxus, and traditionally divided into two groups: the Pleurogona and the Enterogona. An enterogonan ascidian, Ciona intestinalis (Ci), possesses nine Hox genes localized on two chromosomes; thus, the Hox gene cluster is disintegrated. We investigated the Hox gene cluster of a pleurogonan ascidian, Halocynthia roretzi (Hr) to investigate whether Hox gene cluster disintegration is common among ascidians, and if so, how such disintegration occurred during ascidian or tunicate evolution.
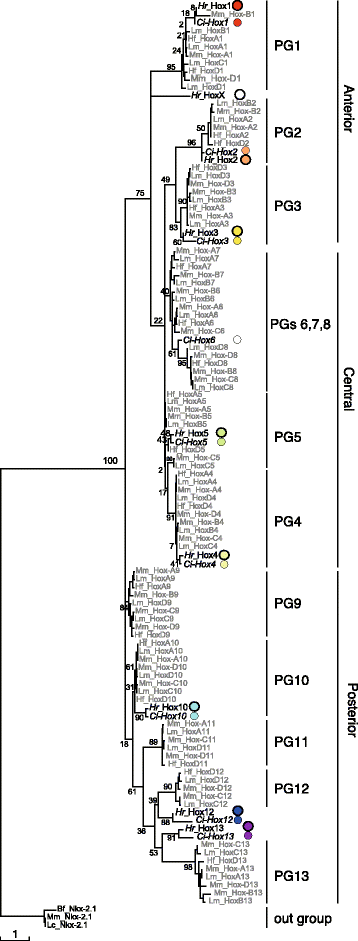
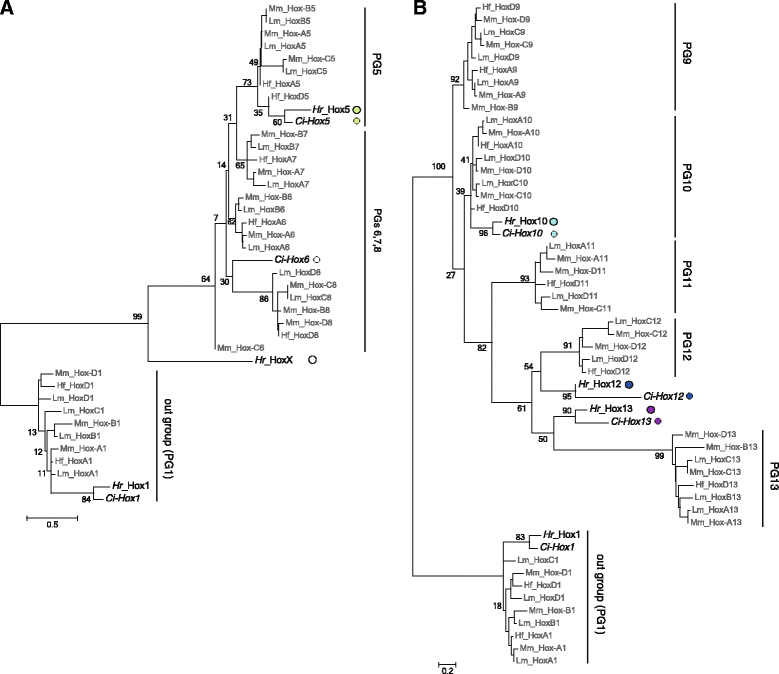
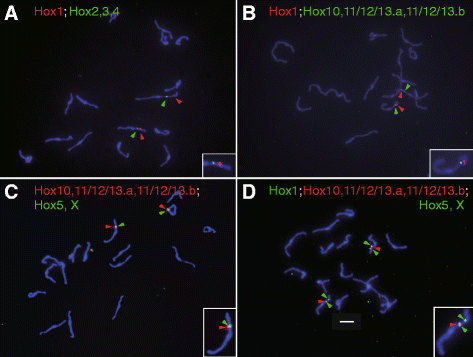
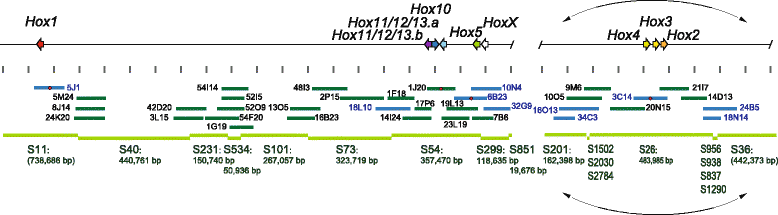
Results: Our phylogenetic analysis reveals that the Hr Hox gene complement comprises nine members, including one with a relatively divergent Hox homeodomain sequence. Eight of nine Hr Hox genes were orthologous to Ci-Hox1, 2, 3, 4, 5, 10, 12 and 13. Following the phylogenetic classification into 13 PGs, we designated Hr Hox genes as Hox1, 2, 3, 4, 5, 10, 11/12/13.a, 11/12/13.b and HoxX. To address the chromosomal arrangement of the nine Hox genes, we performed two-color chromosomal fluorescent in situ hybridization, which revealed that the nine Hox genes are localized on a single chromosome in Hr, distinct from their arrangement in Ci. We further examined the order of the nine Hox genes on the chromosome by chromosome/scaffold walking. This analysis suggested a gene order of Hox1, 11/12/13.b, 11/12/13.a, 10, 5, X, followed by either Hox4, 3, 2 or Hox2, 3, 4 on the chromosome. Based on the present results and those previously reported in Ci, we discuss the establishment of the Hox gene complement and disintegration of Hox gene clusters during the course of ascidian or tunicate evolution.
Conclusions: The Hox gene cluster and the genome must have experienced extensive reorganization during the course of evolution from the ancestral tunicate to Hr and Ci. Nevertheless, some features are shared in Hox gene components and gene arrangement on the chromosomes, suggesting that Hox gene cluster disintegration in ascidians involved early events common to tunicates as well as later ascidian lineage-specific events.
Keywords: Ascidian; Halocynthia roretzi; Hox gene cluster; Tunicate (urochordate) evolution.
Conflict of interest statement
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Figures
Similar articles
-
Note to: Hox gene cluster of the ascidian, Halocynthia roretzi, reveals multiple ancient steps of cluster disintegration during ascidian evolution.Zoological Lett. 2019 Feb 27;5:8. doi: 10.1186/s40851-019-0121-7. eCollection 2019. Zoological Lett. 2019. PMID: 30858988 Free PMC article.
-
The structural organization of ascidian Halocynthia roretzi troponin I genes.J Biochem. 2002 Jul;132(1):135-41. doi: 10.1093/oxfordjournals.jbchem.a003191. J Biochem. 2002. PMID: 12097170
-
Ciona intestinalis Hox gene cluster: Its dispersed structure and residual colinear expression in development.Proc Natl Acad Sci U S A. 2004 Oct 19;101(42):15118-23. doi: 10.1073/pnas.0401389101. Epub 2004 Oct 6. Proc Natl Acad Sci U S A. 2004. PMID: 15469921 Free PMC article.
-
Organization of Hox genes in ascidians: present, past, and future.Dev Dyn. 2005 Jun;233(2):382-9. doi: 10.1002/dvdy.20374. Dev Dyn. 2005. PMID: 15844201 Review.
-
Ascidian actin genes: developmental regulation of gene expression and molecular evolution.Zoolog Sci. 1997 Oct;14(5):707-18. doi: 10.2108/zsj.14.707. Zoolog Sci. 1997. PMID: 9450384 Review.
Cited by
-
The Genome of the "Sea Vomit" Didemnum vexillum.Life (Basel). 2021 Dec 10;11(12):1377. doi: 10.3390/life11121377. Life (Basel). 2021. PMID: 34947908 Free PMC article.
-
Inferring Tunicate Relationships and the Evolution of the Tunicate Hox Cluster with the Genome of Corella inflata.Genome Biol Evol. 2020 Jun 1;12(6):948-964. doi: 10.1093/gbe/evaa060. Genome Biol Evol. 2020. PMID: 32211845 Free PMC article.
-
How Weird is The Worm? Evolution of the Developmental Gene Toolkit in Caenorhabditis elegans.J Dev Biol. 2019 Sep 28;7(4):19. doi: 10.3390/jdb7040019. J Dev Biol. 2019. PMID: 31569401 Free PMC article. Review.
-
The Degenerate Tale of Ascidian Tails.Integr Comp Biol. 2021 Sep 8;61(2):358-369. doi: 10.1093/icb/icab022. Integr Comp Biol. 2021. PMID: 33881514 Free PMC article. Review.
-
Genome-Wide Identification, Comparison, and Expression Analysis of Transcription Factors in Ascidian Styela clava.Int J Mol Sci. 2021 Apr 21;22(9):4317. doi: 10.3390/ijms22094317. Int J Mol Sci. 2021. PMID: 33919240 Free PMC article.
References
LinkOut - more resources
Full Text Sources
Other Literature Sources

