A pathways-based prediction model for classifying breast cancer subtypes
- PMID: 28938599
- PMCID: PMC5601695
- DOI: 10.18632/oncotarget.18544
A pathways-based prediction model for classifying breast cancer subtypes
Abstract
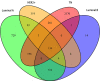
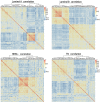
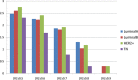
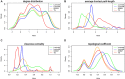
Breast cancer is highly heterogeneous and is classified into four subtypes characterized by specific biological traits, treatment responses, and clinical prognoses. We performed a systemic analysis of 698 breast cancer patient samples from The Cancer Genome Atlas project database. We identified 136 breast cancer genes differentially expressed among the four subtypes. Based on unsupervised clustering analysis, these 136 core genes efficiently categorized breast cancer patients into the appropriate subtypes. Functional enrichment based on Kyoto Encyclopedia of Genes and Genomes analysis identified six functional pathways regulated by these genes: JAK-STAT signaling, basal cell carcinoma, inflammatory mediator regulation of TRP channels, non-small cell lung cancer, glutamatergic synapse, and amyotrophic lateral sclerosis. Three support vector machine (SVM) classification models based on the identified pathways effectively classified different breast cancer subtypes, suggesting that breast cancer subtype-specific risk assessment based on disease pathways could be a potentially valuable approach. Our analysis not only provides insight into breast cancer subtype-specific mechanisms, but also may improve the accuracy of SVM classification models.
Keywords: breast cancer; classification prediction model; co-expression network; pathway enrichment; subtype-specific gene.
Conflict of interest statement
CONFLICTS OF INTEREST The authors declare no conflicts of interest.
Figures

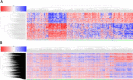

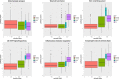

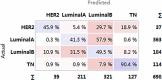

References
-
- Zhu X, Ying J, Wang F, Wang J, Yang H. Estrogen receptor, progesterone receptor, and human epidermal growth factor receptor 2 status in invasive breast cancer: a 3,198 cases study at National Cancer Center, China. Breast Cancer Res Treat. 2014;147:551–555. - PubMed
-
- Prat A, Pineda E, Adamo B, Galván P, Fernández A, Gaba L, Díez M, Viladot M, Arance A, Muñoz M. Clinical implications of the intrinsic molecular subtypes of breast cancer. Breast. 2015;24:S26–35. - PubMed
-
- Chia SK, Speers CH, D’yachkova Y, Kang A, Malfair-Taylor S, Barnett J, Coldman A, Gelmon KA, O’reilly SE, Olivotto IA. The impact of new chemotherapeutic and hormone agents on survival in a population-based cohort of women with metastatic breast cancer. Cancer. 2007;110:973–9. - PubMed
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

