Identification of potential cancer-related pseudogenes in lung adenocarcinoma based on ceRNA hypothesis
- PMID: 28938616
- PMCID: PMC5601712
- DOI: 10.18632/oncotarget.19933
Identification of potential cancer-related pseudogenes in lung adenocarcinoma based on ceRNA hypothesis
Abstract
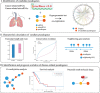
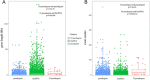
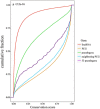
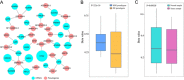
Pseudogenes are initially regarded as non-functional genomic fossils resulted from inactivating gene mutations during evolution. Far from being silent, pseudogenes are proved to regulate the expression of protein-coding genes through function as microRNA sponge in vivo. The aim of our study was to propose an integrative systems biology approach to identify disease pseudogenes base on competitive endogenous RNA (ceRNA) hypothesis. Here, we applied our method to lung adenocarcinoma (LUAD) RNASeq data from TCGA and identified 33 candidate pseudogenes. We described the characteristics of the candidate pseudogenes and performed functional enrichment. Through analyzing neighboring genes we found these pseudogenes were surrounded by tumor genes and may involve in tumor pathway. Furthermore, the DNA methylation analysis indicated that 21 pseudogenes co-methylated with their competitive mRNAs. In the co-methylated network, we discovered 6 differentially expressed pseudogenes, which we termed potential LUAD-associated pseudogenes. We further revealed that the 3 ceRNA triples (miR-21-5p-NKAPP1-PRDM11, miR-29c-3p-MSTO2P-EZH2 and miR-29c-3p-RPLP0P2-EZH2), whose high risk groups were associated with the poor prognosis of LUAD, may be considered as potential prognostic signatures. Moreover, by integrating target information of microRNA we also provided a new perspective for the discovery of potential small molecule drugs. This work may facilitate cancer research and serve as the basis for future efforts to understand the role of pseudogenes, develop novel biomarkers and improve knowledge of tumor biology.
Keywords: ceRNA; lung adenocarcinoma; pseudogenes.
Conflict of interest statement
CONFLICTS OF INTEREST The authors declared that they have no conflicts of interest.
Figures

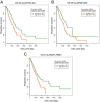

References
-
- Watanabe T, Totoki Y, Toyoda A, Kaneda M, Kuramochi-Miyagawa S, Obata Y, Chiba H, Kohara Y, Kono T, Nakano T, Surani MA, Sakaki Y, Sasaki H. Endogenous siRNAs from naturally formed dsRNAs regulate transcripts in mouse oocytes. Nature. 2008;453:539–543. - PubMed
-
- Poliseno L. Pseudogenes: newly discovered players in human cancer. Sci Signal. 2012;5:re5. - PubMed
LinkOut - more resources
Full Text Sources
Other Literature Sources

