Experimental Evolution of Diverse Strains as a Method for the Determination of Biochemical Mechanisms of Action for Novel Pyrrolizidinone Antibiotics
- PMID: 28942642
- PMCID: PMC5817888
- DOI: 10.1021/acsinfecdis.7b00135
Experimental Evolution of Diverse Strains as a Method for the Determination of Biochemical Mechanisms of Action for Novel Pyrrolizidinone Antibiotics
Abstract
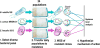
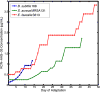
The continuing rise of multidrug resistant pathogens has made it clear that in the absence of new antibiotics we are moving toward a "postantibiotic" world, in which even routine infections will become increasingly untreatable. There is a clear need for the development of new antibiotics with truly novel mechanisms of action to combat multidrug resistant pathogens. Experimental evolution to resistance can be a useful tactic for the characterization of the biochemical mechanism of action for antibiotics of interest. Herein, we demonstrate that the use of a diverse panel of strains with well-annotated reference genomes improves the success of using experimental evolution to characterize the mechanism of action of a novel pyrrolizidinone antibiotic analog. Importantly, we used experimental evolution under conditions that favor strongly polymorphic populations to adapt a panel of three substantially different Gram-positive species (lab strain Bacillus subtilis and clinical strains methicillin-resistant Staphylococcus aureus MRSA131 and Enterococcus faecalis S613) to produce a sufficiently diverse set of evolutionary outcomes. Comparative whole genome sequencing (WGS) between the susceptible starting strain and the resistant strains was then used to identify the genetic changes within each species in response to the pyrrolizidinone. Taken together, the adaptive response across a range of organisms allowed us to develop a readily testable hypothesis for the mechanism of action of the CJ-16 264 analog. In conjunction with mitochondrial inhibition studies, we were able to elucidate that this novel pyrrolizidinone antibiotic is an electron transport chain (ETC) inhibitor. By studying evolution to resistance in a panel of different species of bacteria, we have developed an enhanced method for the characterization of new lead compounds for the discovery of new mechanisms of action.
Keywords: antibiotic resistance; antibiotics; experimental evolution; mechanism of action; pyrrolizidinone.
Conflict of interest statement
The authors declare no competing financial interest.
Figures



Similar articles
-
Using experimental evolution to identify druggable targets that could inhibit the evolution of antimicrobial resistance.J Antibiot (Tokyo). 2018 Feb;71(2):279-286. doi: 10.1038/ja.2017.108. Epub 2017 Sep 20. J Antibiot (Tokyo). 2018. PMID: 28928474 Free PMC article.
-
Synergistic activity profile of an antimicrobial peptide against multidrug-resistant and extensively drug-resistant strains of Gram-negative bacterial pathogens.J Pept Sci. 2017 Apr;23(4):329-333. doi: 10.1002/psc.2978. Epub 2017 Feb 8. J Pept Sci. 2017. PMID: 28176481
-
Isolation, structure elucidation and antibacterial activity of a new tetramic acid, ascosetin.J Antibiot (Tokyo). 2014 Jul;67(7):527-31. doi: 10.1038/ja.2014.33. Epub 2014 Apr 2. J Antibiot (Tokyo). 2014. PMID: 24690911
-
Multidrug-Resistant Enterococcal Infections: New Compounds, Novel Antimicrobial Therapies?Trends Microbiol. 2017 Jun;25(6):467-479. doi: 10.1016/j.tim.2017.01.004. Epub 2017 Feb 13. Trends Microbiol. 2017. PMID: 28209400 Review.
-
Development of novel antibacterial drugs to combat multiple resistant organisms.Langenbecks Arch Surg. 2015 Feb;400(2):153-65. doi: 10.1007/s00423-015-1280-4. Epub 2015 Feb 11. Langenbecks Arch Surg. 2015. PMID: 25667169 Review.
Cited by
-
Microfluidic platform for spatially segregated experimental evolution studies with E. coli.STAR Protoc. 2022 Apr 21;3(2):101332. doi: 10.1016/j.xpro.2022.101332. eCollection 2022 Jun 17. STAR Protoc. 2022. PMID: 35496805 Free PMC article.
-
Novel Immune Modulators Enhance Caenorhabditis elegans Resistance to Multiple Pathogens.mSphere. 2021 Jan 6;6(1):e00950-20. doi: 10.1128/mSphere.00950-20. mSphere. 2021. PMID: 33408224 Free PMC article.
-
Enantioselective Total Synthesis of Antibiotic CJ-16,264, Synthesis and Biological Evaluation of Designed Analogues, and Discovery of Highly Potent and Simpler Antibacterial Agents.J Am Chem Soc. 2017 Nov 8;139(44):15868-15877. doi: 10.1021/jacs.7b08749. Epub 2017 Oct 24. J Am Chem Soc. 2017. PMID: 29064688 Free PMC article.
-
Identification of Evolutionary Trajectories Associated with Antimicrobial Resistance Using Microfluidics.ACS Infect Dis. 2022 Jan 14;8(1):242-254. doi: 10.1021/acsinfecdis.1c00564. Epub 2021 Dec 28. ACS Infect Dis. 2022. PMID: 34962128 Free PMC article.
-
Repurposing bioactive compounds for treating multidrug-resistant pathogens.J Med Microbiol. 2020 Jun;69(6):881-894. doi: 10.1099/jmm.0.001172. J Med Microbiol. 2020. PMID: 32163353 Free PMC article.
References
-
- Blair JMA, Webber MA, Baylay AJ, Ogbolu DO, Piddock LJV. Molecular mechanisms of antibiotic resistance. Nat Rev Microbiol. 2015;13(1):42–51. - PubMed
-
- CDC. Antibiotic resistance threats in the United States, 2013. CDC; Atlanta, GA: 2013.
-
- The PEW Charitable Trusts. Antibiotics Currently in Clinical Development. 2016 http://www.pewtrusts.org/~/media/assets/2016/05/antibiotics-currently-in....
-
- Banerjee A, Dubnau E, Quemard A, Balasubramanian V, Um KS, Wilson T, Collins D, de Lisle G, Jacobs WR. inhA, a gene encoding a target for isoniazid and ethionamide in Mycobacterium tuberculosis. Science. 1994;263(5144):227–230. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Miscellaneous

