Benchmarking viromics: an in silico evaluation of metagenome-enabled estimates of viral community composition and diversity
- PMID: 28948103
- PMCID: PMC5610896
- DOI: 10.7717/peerj.3817
Benchmarking viromics: an in silico evaluation of metagenome-enabled estimates of viral community composition and diversity
Abstract
Background: Viral metagenomics (viromics) is increasingly used to obtain uncultivated viral genomes, evaluate community diversity, and assess ecological hypotheses. While viromic experimental methods are relatively mature and widely accepted by the research community, robust bioinformatics standards remain to be established. Here we used in silico mock viral communities to evaluate the viromic sequence-to-ecological-inference pipeline, including (i) read pre-processing and metagenome assembly, (ii) thresholds applied to estimate viral relative abundances based on read mapping to assembled contigs, and (iii) normalization methods applied to the matrix of viral relative abundances for alpha and beta diversity estimates.
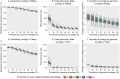
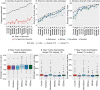
Results: Tools specifically designed for metagenomes, specifically metaSPAdes, MEGAHIT, and IDBA-UD, were the most effective at assembling viromes. Read pre-processing, such as partitioning, had virtually no impact on assembly output, but may be useful when hardware is limited. Viral populations with 2-5 × coverage typically assembled well, whereas lesser coverage led to fragmented assembly. Strain heterogeneity within populations hampered assembly, especially when strains were closely related (average nucleotide identity, or ANI ≥97%) and when the most abundant strain represented <50% of the population. Viral community composition assessments based on read recruitment were generally accurate when the following thresholds for detection were applied: (i) ≥10 kb contig lengths to define populations, (ii) coverage defined from reads mapping at ≥90% identity, and (iii) ≥75% of contig length with ≥1 × coverage. Finally, although data are limited to the most abundant viruses in a community, alpha and beta diversity patterns were robustly estimated (±10%) when comparing samples of similar sequencing depth, but more divergent (up to 80%) when sequencing depth was uneven across the dataset. In the latter cases, the use of normalization methods specifically developed for metagenomes provided the best estimates.
Conclusions: These simulations provide benchmarks for selecting analysis cut-offs and establish that an optimized sample-to-ecological-inference viromics pipeline is robust for making ecological inferences from natural viral communities. Continued development to better accessing RNA, rare, and/or diverse viral populations and improved reference viral genome availability will alleviate many of viromics remaining limitations.
Keywords: Assembly; Benchmarks; Metagenome; Viral ecology; Virome; Virus.
Conflict of interest statement
The authors declare there are no competing interests.
Figures


Similar articles
-
Expanding standards in viromics: in silico evaluation of dsDNA viral genome identification, classification, and auxiliary metabolic gene curation.PeerJ. 2021 Jun 14;9:e11447. doi: 10.7717/peerj.11447. eCollection 2021. PeerJ. 2021. PMID: 34178438 Free PMC article.
-
Fragmentation and Coverage Variation in Viral Metagenome Assemblies, and Their Effect in Diversity Calculations.Front Bioeng Biotechnol. 2015 Sep 17;3:141. doi: 10.3389/fbioe.2015.00141. eCollection 2015. Front Bioeng Biotechnol. 2015. PMID: 26442255 Free PMC article.
-
MVP: a modular viromics pipeline to identify, filter, cluster, annotate, and bin viruses from metagenomes.mSystems. 2024 Oct 22;9(10):e0088824. doi: 10.1128/msystems.00888-24. Epub 2024 Oct 1. mSystems. 2024. PMID: 39352141 Free PMC article.
-
Assessment of metagenomic assemblers based on hybrid reads of real and simulated metagenomic sequences.Brief Bioinform. 2020 May 21;21(3):777-790. doi: 10.1093/bib/bbz025. Brief Bioinform. 2020. PMID: 30860572 Free PMC article. Review.
-
Recovering complete and draft population genomes from metagenome datasets.Microbiome. 2016 Mar 8;4:8. doi: 10.1186/s40168-016-0154-5. Microbiome. 2016. PMID: 26951112 Free PMC article. Review.
Cited by
-
Metagenomic Data Assembly - The Way of Decoding Unknown Microorganisms.Front Microbiol. 2021 Mar 23;12:613791. doi: 10.3389/fmicb.2021.613791. eCollection 2021. Front Microbiol. 2021. PMID: 33833738 Free PMC article. Review.
-
Virome assembly and annotation in brain tissue based on next-generation sequencing.Cancer Med. 2020 Sep;9(18):6776-6790. doi: 10.1002/cam4.3325. Epub 2020 Aug 1. Cancer Med. 2020. PMID: 32738030 Free PMC article.
-
Efficient Recovery of Complete Gut Viral Genomes by Combined Short- and Long-Read Sequencing.Adv Sci (Weinh). 2024 Apr;11(13):e2305818. doi: 10.1002/advs.202305818. Epub 2024 Jan 19. Adv Sci (Weinh). 2024. PMID: 38240578 Free PMC article.
-
Freshwater Mussels Show Elevated Viral Richness and Intensity during a Mortality Event.Viruses. 2022 Nov 23;14(12):2603. doi: 10.3390/v14122603. Viruses. 2022. PMID: 36560607 Free PMC article.
-
Benchmarking protocols for the metagenomic analysis of stream biofilm viromes.PeerJ. 2019 Dec 20;7:e8187. doi: 10.7717/peerj.8187. eCollection 2019. PeerJ. 2019. PMID: 31879573 Free PMC article.
References
-
- Allers E, Moraru C, Duhaime MB, Beneze E, Solonenko N, Canosa JB, Amann R, Sullivan MB. Single-cell and population level viral infection dynamics revealed by phageFISH, a method to visualize intracellular and free viruses. Environmental Microbiology. 2013;15:2306–2318. doi: 10.1111/1462-2920.12100. - DOI - PMC - PubMed
-
- Angly FE, Willner D, Prieto-Davó A, Edwards RA, Schmieder R, Vega-Thurber R, Antonopoulos DA, Barott K, Cottrell MT, Desnues C, Dinsdale EA, Furlan M, Haynes M, Henn MR, Hu Y, Kirchman DL, McDole T, McPherson JD, Meyer F, Miller RM, Mundt E, Naviaux RK, Rodriguez-Mueller B, Stevens R, Wegley L, Zhang L, Zhu B, Rohwer F. The GAAS metagenomic tool and its estimations of viral and microbial average genome size in four major biomes. PLOS Computational Biology. 2009;5:e1000593. doi: 10.1371/journal.pcbi.1000593. - DOI - PMC - PubMed
LinkOut - more resources
Full Text Sources
Other Literature Sources

