CG dinucleotide suppression enables antiviral defence targeting non-self RNA
- PMID: 28953888
- PMCID: PMC6592701
- DOI: 10.1038/nature24039
CG dinucleotide suppression enables antiviral defence targeting non-self RNA
Abstract
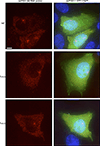
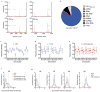
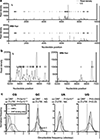
Vertebrate genomes exhibit marked CG suppression-that is, lower than expected numbers of 5'-CG-3' dinucleotides. This feature is likely to be due to C-to-T mutations that have accumulated over hundreds of millions of years, driven by CG-specific DNA methyl transferases and spontaneous methyl-cytosine deamination. Many RNA viruses of vertebrates that are not substrates for DNA methyl transferases mimic the CG suppression of their hosts. This property of viral genomes is unexplained. Here we show, using synonymous mutagenesis, that CG suppression is essential for HIV-1 replication. The deleterious effect of CG dinucleotides on HIV-1 replication was cumulative, associated with cytoplasmic RNA depletion, and was exerted by CG dinucleotides in both translated and non-translated exonic RNA sequences. A focused screen using small inhibitory RNAs revealed that zinc-finger antiviral protein (ZAP) inhibited virion production by cells infected with CG-enriched HIV-1. Crucially, HIV-1 mutants containing segments whose CG content mimicked random nucleotide sequence were defective in unmanipulated cells, but replicated normally in ZAP-deficient cells. Crosslinking-immunoprecipitation-sequencing assays demonstrated that ZAP binds directly and selectively to RNA sequences containing CG dinucleotides. These findings suggest that ZAP exploits host CG suppression to identify non-self RNA. The dinucleotide composition of HIV-1, and perhaps other RNA viruses, appears to have adapted to evade this host defence.
Conflict of interest statement
The Authors declare that they have no competing financial interests.
Figures









Comment in
-
Evolution: Zapping viral RNAs.Nature. 2017 Oct 5;550(7674):46-47. doi: 10.1038/nature24140. Epub 2017 Sep 27. Nature. 2017. PMID: 28953872 Free PMC article.
-
Viral infection: Adapt or get zapped.Nat Rev Microbiol. 2017 Oct 12;15(11):641. doi: 10.1038/nrmicro.2017.129. Nat Rev Microbiol. 2017. PMID: 29021599 No abstract available.
-
Mind Your Cs and Gs.Cell Host Microbe. 2017 Nov 8;22(5):575-576. doi: 10.1016/j.chom.2017.10.012. Cell Host Microbe. 2017. PMID: 29120736
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

