Molecular Analysis of a Short-term Model of β-Glucans-Trained Immunity Highlights the Accessory Contribution of GM-CSF in Priming Mouse Macrophages Response
- PMID: 28955331
- PMCID: PMC5601002
- DOI: 10.3389/fimmu.2017.01089
Molecular Analysis of a Short-term Model of β-Glucans-Trained Immunity Highlights the Accessory Contribution of GM-CSF in Priming Mouse Macrophages Response
Abstract
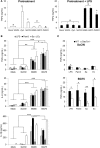
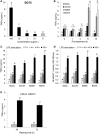
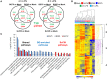
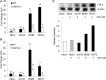
β-Glucans (BGs) are glucose polymers present in the fungal cell wall (CW) and, as such, are recognized by innate immune cells as microbial-associated pattern through Dectin-1 receptor. Recent studies have highlighted the ability of the pathogenic yeast Candida albicans or its CW-derived β(1,3) (1,6)-glucans to increase human monocytes cytokine secretion upon secondary stimulation, a phenomenon now referred as immune training. This ability of monocytes programming confers BGs an undeniable immunotherapeutic potential. Our objective was to determine whether BGs from Saccharomyces cerevisiae, a non-pathogenic yeast, are endowed with such a property. For this purpose, we have developed a short-term training model based on lipopolysaccharide re-stimulation of mouse bone marrow-derived macrophages primed with S. cerevisiae BGs. Through a transcriptome analysis, we demonstrated that BGs induced a specific gene expression signature involving the PI3K/AKT signaling pathway as in human monocytes. Moreover, we showed that over-expression of Csf2 (that encodes for GM-CSF) was a Dectin-1-dependent feature of BG-induced priming of macrophages. Further experiments confirmed that GM-CSF up-regulated Dectin-1 cell surface expression and amplified macrophages response along BG-mediated training. However, the blockade of GM-CSFR demonstrated that GM-CSF was not primarily required for BG-induced training of macrophages although it can substantially improve it. In addition, we found that mouse macrophages trained with BGs upregulated their expression of the four and a half LIM-only protein 2 (Fhl2) in a Dectin-1-dependent manner. Consistently, we observed that intracellular levels of FHL2 increased after stimulation of macrophages with BGs. In conclusion, our experiments provide new insights on GM-CSF contribution to the training of cells from the monocytic lineage and highlights FHL2 as a possible regulator of BG-associated signaling.
Keywords: Dectin-1; GM-CSF; macrophage; trained immunity; β-glucans.
Figures

Similar articles
-
Soluble β-glucan from Grifola frondosa induces proliferation and Dectin-1/Syk signaling in resident macrophages via the GM-CSF autocrine pathway.J Leukoc Biol. 2012 Apr;91(4):547-56. doi: 10.1189/jlb.0711386. Epub 2011 Oct 25. J Leukoc Biol. 2012. PMID: 22028332
-
(1,3)-beta-glucans activate both dectin-1 and NLRP3 inflammasome in human macrophages.J Immunol. 2010 Jun 1;184(11):6335-42. doi: 10.4049/jimmunol.0903019. Epub 2010 Apr 26. J Immunol. 2010. PMID: 20421639
-
More Prominent Inflammatory Response to Pachyman than to Whole-Glucan Particle and Oat-β-Glucans in Dextran Sulfate-Induced Mucositis Mice and Mouse Injection through Proinflammatory Macrophages.Int J Mol Sci. 2022 Apr 5;23(7):4026. doi: 10.3390/ijms23074026. Int J Mol Sci. 2022. PMID: 35409384 Free PMC article.
-
Contribution of dectin-1 and granulocyte macrophage-colony stimulating factor (GM-CSF) to immunomodulating actions of beta-glucan.Int Immunopharmacol. 2008 Apr;8(4):556-66. doi: 10.1016/j.intimp.2007.12.011. Epub 2008 Jan 16. Int Immunopharmacol. 2008. PMID: 18328447 Review.
-
Long-lived effects of administering β-glucans: Indications for trained immunity in fish.Dev Comp Immunol. 2016 Nov;64:93-102. doi: 10.1016/j.dci.2016.03.003. Epub 2016 Mar 2. Dev Comp Immunol. 2016. PMID: 26945622 Review.
Cited by
-
Effects of dietary β-glucan and rice fermented on growth performance, fatty acids, and Newcastle disease immune response in turkey broilers.Saudi J Biol Sci. 2023 Aug;30(8):103736. doi: 10.1016/j.sjbs.2023.103736. Epub 2023 Jul 13. Saudi J Biol Sci. 2023. PMID: 37521751 Free PMC article.
-
Trained Immunity as a Prospective Tool against Emerging Respiratory Pathogens.Vaccines (Basel). 2022 Nov 15;10(11):1932. doi: 10.3390/vaccines10111932. Vaccines (Basel). 2022. PMID: 36423027 Free PMC article. Review.
-
Analysis of the Long-Lived Responses Induced by Immunostimulants and Their Effects on a Viral Infection in Zebrafish (Danio rerio).Front Immunol. 2018 Jul 9;9:1575. doi: 10.3389/fimmu.2018.01575. eCollection 2018. Front Immunol. 2018. PMID: 30038625 Free PMC article.
-
Targeting SHIP-1 in Myeloid Cells Enhances Trained Immunity and Boosts Response to Infection.Cell Rep. 2018 Oct 30;25(5):1118-1126. doi: 10.1016/j.celrep.2018.09.092. Cell Rep. 2018. PMID: 30380404 Free PMC article.
-
Fungal dysbiosis of the gut microbiota is associated with colorectal cancer in Chinese patients.Am J Transl Res. 2021 Oct 15;13(10):11287-11301. eCollection 2021. Am J Transl Res. 2021. PMID: 34786058 Free PMC article.
References
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

