Challenges for Super-Resolution Localization Microscopy and Biomolecular Fluorescent Nano-Probing in Cancer Research
- PMID: 28956810
- PMCID: PMC5666748
- DOI: 10.3390/ijms18102066
Challenges for Super-Resolution Localization Microscopy and Biomolecular Fluorescent Nano-Probing in Cancer Research
Abstract
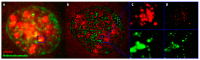
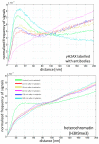
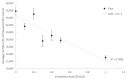
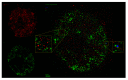
Understanding molecular interactions and regulatory mechanisms in tumor initiation, progression, and treatment response are key requirements towards advanced cancer diagnosis and novel treatment procedures in personalized medicine. Beyond decoding the gene expression, malfunctioning and cancer-related epigenetic pathways, investigations of the spatial receptor arrangements in membranes and genome organization in cell nuclei, on the nano-scale, contribute to elucidating complex molecular mechanisms in cells and tissues. By these means, the correlation between cell function and spatial organization of molecules or molecular complexes can be studied, with respect to carcinogenesis, tumor sensitivity or tumor resistance to anticancer therapies, like radiation or antibody treatment. Here, we present several new applications for bio-molecular nano-probes and super-resolution, laser fluorescence localization microscopy and their potential in life sciences, especially in biomedical and cancer research. By means of a tool-box of fluorescent antibodies, green fluorescent protein (GFP) tagging, or specific oligonucleotides, we present tumor relevant re-arrangements of Erb-receptors in membranes, spatial organization of Smad specific ubiquitin protein ligase 2 (Smurf2) in the cytosol, tumor cell characteristic heterochromatin organization, and molecular re-arrangements induced by radiation or antibody treatment. The main purpose of this article is to demonstrate how nano-scaled distance measurements between bio-molecules, tagged by appropriate nano-probes, can be applied to elucidate structures and conformations of molecular complexes which are characteristic of tumorigenesis and treatment responses. These applications open new avenues towards a better interpretation of the spatial organization and treatment responses of functionally relevant molecules, at the single cell level, in normal and cancer cells, offering new potentials for individualized medicine.
Keywords: Smurf2; cancer research; chromatin organization; chromatin radiation response; fluorescent nano-probes; receptor conformation changes; super-resolution localization microscopy; γ-H2AX phosphorylation sites.
Conflict of interest statement
The authors declare no conflict of interest. The founding sponsors had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the manuscript, and in the decision to publish the results.
Figures
Similar articles
-
Combining Low Temperature Fluorescence DNA-Hybridization, Immunostaining, and Super-Resolution Localization Microscopy for Nano-Structure Analysis of ALU Elements and Their Influence on Chromatin Structure.Int J Mol Sci. 2017 May 7;18(5):1005. doi: 10.3390/ijms18051005. Int J Mol Sci. 2017. PMID: 28481278 Free PMC article.
-
Spatial Arrangements of Connexin43 in Cancer Related Cells and Re-Arrangements under Treatment Conditions: Investigations on the Nano-Scale by Super-Resolution Localization Light Microscopy.Cancers (Basel). 2019 Mar 4;11(3):301. doi: 10.3390/cancers11030301. Cancers (Basel). 2019. PMID: 30836676 Free PMC article.
-
In situ optical sequencing and structure analysis of a trinucleotide repeat genome region by localization microscopy after specific COMBO-FISH nano-probing.Nanoscale. 2015 Nov 14;7(42):17938-46. doi: 10.1039/c5nr04141d. Epub 2015 Oct 14. Nanoscale. 2015. PMID: 26463479
-
Super-resolution microscopy approaches to nuclear nanostructure imaging.Methods. 2017 Jul 1;123:11-32. doi: 10.1016/j.ymeth.2017.03.019. Epub 2017 Apr 6. Methods. 2017. PMID: 28390838 Review.
-
DNA and chromatin imaging with super-resolution fluorescence microscopy based on single-molecule localization.Biopolymers. 2011 May;95(5):290-7. doi: 10.1002/bip.21574. Epub 2010 Dec 23. Biopolymers. 2011. PMID: 21184489 Review.
Cited by
-
Incorporation of Low Concentrations of Gold Nanoparticles: Complex Effects on Radiation Response and Fate of Cancer Cells.Pharmaceutics. 2022 Jan 11;14(1):166. doi: 10.3390/pharmaceutics14010166. Pharmaceutics. 2022. PMID: 35057061 Free PMC article.
-
Visualizing ubiquitination in mammalian cells.EMBO Rep. 2019 Feb;20(2):e46520. doi: 10.15252/embr.201846520. Epub 2019 Jan 21. EMBO Rep. 2019. PMID: 30665942 Free PMC article. Review.
-
A Paradigm Revolution or Just Better Resolution-Will Newly Emerging Superresolution Techniques Identify Chromatin Architecture as a Key Factor in Radiation-Induced DNA Damage and Repair Regulation?Cancers (Basel). 2020 Dec 23;13(1):18. doi: 10.3390/cancers13010018. Cancers (Basel). 2020. PMID: 33374540 Free PMC article. Review.
-
A comprehensive method to study the DNA's association with lamin and chromatin compaction in intact cell nuclei at super resolution.Nanoscale. 2023 Jan 5;15(2):742-756. doi: 10.1039/d2nr02684h. Nanoscale. 2023. PMID: 36524744 Free PMC article.
-
From survival of irradiated mice to modern molecular insights: a seventy-year journey in radiobiology at the institute of biophysics, Czech academy of sciences.Eur Biophys J. 2025 Jun 13. doi: 10.1007/s00249-025-01765-9. Online ahead of print. Eur Biophys J. 2025. PMID: 40512188 Review.
References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

