Covalent Lectin Inhibition and Application in Bacterial Biofilm Imaging
- PMID: 28960731
- PMCID: PMC5767747
- DOI: 10.1002/anie.201709368
Covalent Lectin Inhibition and Application in Bacterial Biofilm Imaging
Abstract
Biofilm formation by pathogenic bacteria is a hallmark of chronic infections. In many cases, lectins play key roles in establishing biofilms. The pathogen Pseudomonas aeruginosa often exhibiting various drug resistances employs its lectins LecA and LecB as virulence factors and biofilm building blocks. Therefore, inhibition of the function of these proteins is thought to have potential in developing "pathoblockers" preventing biofilm formation and virulence. A covalent lectin inhibitor specific to a carbohydrate binding site is described for the first time. Its application in the LecA-specific in vitro imaging of biofilms formed by P. aeruginosa is also reported.
Keywords: Pseudomonas aeruginosa; biofilms; carbohydrates; glycomimetics; lectins.
© 2017 The Authors. Published by Wiley-VCH Verlag GmbH & Co. KGaA.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

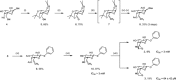
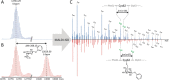





References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

