Characterization of HIV diversity, phylodynamics and drug resistance in Washington, DC
- PMID: 28961263
- PMCID: PMC5621693
- DOI: 10.1371/journal.pone.0185644
Characterization of HIV diversity, phylodynamics and drug resistance in Washington, DC
Abstract
Background: Washington DC has a high burden of HIV with a 2.0% HIV prevalence. The city is a national and international hub potentially containing a broad diversity of HIV variants; yet few sequences from DC are available on GenBank to assess the evolutionary history of HIV in the US capital. Towards this general goal, here we analyze extensive sequence data and investigate HIV diversity, phylodynamics, and drug resistant mutations (DRM) in DC.
Methods: Molecular HIV-1 sequences were collected from participants infected through 2015 as part of the DC Cohort, a longitudinal observational study of HIV+ patients receiving care at 13 DC clinics. Sequences were paired with Cohort demographic, risk, and clinical data and analyzed using maximum likelihood, Bayesian and coalescent approaches of phylogenetic, network and population genetic inference. We analyzed 601 sequences from 223 participants for int (~864 bp) and 2,810 sequences from 1,659 participants for PR/RT (~1497 bp).
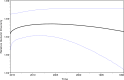
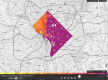
Results: Ninety-nine and 94% of the int and PR/RT sequences, respectively, were identified as subtype B, with 14 non-B subtypes also detected. Phylodynamic analyses of US born infected individuals showed that HIV population size varied little over time with no significant decline in diversity. Phylogenetic analyses grouped 13.5% of the int sequences into 14 clusters of 2 or 3 sequences, and 39.0% of the PR/RT sequences into 203 clusters of 2-32 sequences. Network analyses grouped 3.6% of the int sequences into 4 clusters of 2 sequences, and 10.6% of the PR/RT sequences into 76 clusters of 2-7 sequences. All network clusters were detected in our phylogenetic analyses. Higher proportions of clustered sequences were found in zip codes where HIV prevalence is highest (r = 0.607; P<0.00001). We detected a high prevalence of DRM for both int (17.1%) and PR/RT (39.1%), but only 8 int and 12 PR/RT amino acids were identified as under adaptive selection. We observed a significant (P<0.0001) association between main risk factors (men who have sex with men and heterosexuals) and genotypes in the five well-supported clusters with sufficient sample size for testing.
Discussion: Pairing molecular data with clinical and demographic data provided novel insights into HIV population dynamics in Washington, DC. Identification of populations and geographic locations where clustering occurs can inform and complement active surveillance efforts to interrupt HIV transmission.
Conflict of interest statement
Figures


References
-
- District of Columbia Department of Health HIV/AIDS H, STD, and TB Administration (HAHSTA). Annual Epidemiology & Surveillance Report: Surveillance Data Through December 2016 Washington, DC: DC Department of Health, 2016.
-
- Castel AD, Kalmin MM, Hart RL, Young HA, Hays H, Benator D, et al. Disparities in achieving and sustaining viral suppression among a large cohort of HIV-infected persons in care—Washington, DC. AIDS Care. 2016;28(11):1355–64. doi: 10.1080/09540121.2016.1189496 ; PubMed Central PMCID: PMCPMC5084086. - DOI - PMC - PubMed
-
- Quinn TC, Wawer MJ, Sewankambo N, Serwadda D, Li C, Wabwire-Mangen F, et al. Viral load and heterosexual transmission of human immunodeficiency virus type 1. Rakai Project Study Group. The New England journal of medicine. 2000;342(13):921–9. doi: 10.1056/NEJM200003303421303 . - DOI - PubMed
-
- Counsil AI. New Americans in Washington, D.C.: The Political and Economic Power of Immigrants, Latinos, and Asians in Our Nation’s Capital 2015 [cited 2016 May 6]. Available from: http://www.immigrationpolicy.org/sites/default/files/docs/new_americans_...).
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials

