RNA Interference Screening Reveals Host CaMK4 as a Regulator of Cryptococcal Uptake and Pathogenesis
- PMID: 28970273
- PMCID: PMC5695120
- DOI: 10.1128/IAI.00195-17
RNA Interference Screening Reveals Host CaMK4 as a Regulator of Cryptococcal Uptake and Pathogenesis
Abstract
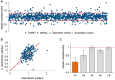
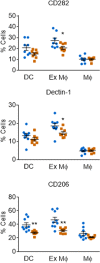
Cryptococcus neoformans, the causative agent of cryptococcosis, is an opportunistic fungal pathogen that kills over 200,000 individuals annually. This yeast may grow freely in body fluids, but it also flourishes within host cells. Despite extensive research on cryptococcal pathogenesis, host genes involved in the initial engulfment of fungi and subsequent stages of infection are woefully understudied. To address this issue, we combined short interfering RNA silencing and a high-throughput imaging assay to identify host regulators that specifically influence cryptococcal uptake. Of 868 phosphatase and kinase genes assayed, we discovered 79 whose silencing significantly affected cryptococcal engulfment. For 25 of these, the effects were fungus specific, as opposed to general alterations in phagocytosis. Four members of this group significantly and specifically altered cryptococcal uptake; one of them encoded CaMK4, a calcium/calmodulin-dependent protein kinase. Pharmacological inhibition of CaMK4 recapitulated the observed defects in phagocytosis. Furthermore, mice deficient in CaMK4 showed increased survival compared to wild-type mice upon infection with C. neoformans This increase in survival correlated with decreased expression of pattern recognition receptors on host phagocytes known to recognize C. neoformans Altogether, we have identified a kinase that is involved in C. neoformans internalization by host cells and in host resistance to this deadly infection.
Keywords: Cryptococcus neoformans; fungal pathogenesis; image-based screen.
Copyright © 2017 American Society for Microbiology.
Figures


Similar articles
-
Cdk8 and Ssn801 Regulate Oxidative Stress Resistance and Virulence in Cryptococcus neoformans.mBio. 2019 Feb 12;10(1):e02818-18. doi: 10.1128/mBio.02818-18. mBio. 2019. PMID: 30755515 Free PMC article.
-
Live Imaging of Host-Parasite Interactions in a Zebrafish Infection Model Reveals Cryptococcal Determinants of Virulence and Central Nervous System Invasion.mBio. 2015 Sep 29;6(5):e01425-15. doi: 10.1128/mBio.01425-15. mBio. 2015. PMID: 26419880 Free PMC article.
-
A Single Protein S-acyl Transferase Acts through Diverse Substrates to Determine Cryptococcal Morphology, Stress Tolerance, and Pathogenic Outcome.PLoS Pathog. 2015 May 13;11(5):e1004908. doi: 10.1371/journal.ppat.1004908. eCollection 2015 May. PLoS Pathog. 2015. PMID: 25970403 Free PMC article.
-
Mechanisms of cryptococcal virulence and persistence.Future Microbiol. 2010 Aug;5(8):1269-88. doi: 10.2217/fmb.10.93. Future Microbiol. 2010. PMID: 20722603 Review.
-
Catch me if you can: phagocytosis and killing avoidance by Cryptococcus neoformans.FEMS Immunol Med Microbiol. 2012 Mar;64(2):147-61. doi: 10.1111/j.1574-695X.2011.00871.x. FEMS Immunol Med Microbiol. 2012. PMID: 22029633 Review.
Cited by
-
An Automated Assay to Measure Phagocytosis of Cryptococcus neoformans.Curr Protoc Microbiol. 2019 Jun;53(1):e79. doi: 10.1002/cpmc.79. Epub 2019 Feb 25. Curr Protoc Microbiol. 2019. PMID: 30802005 Free PMC article.
-
Sac1 links phosphoinositide turnover to cryptococcal virulence.mBio. 2024 Aug 14;15(8):e0149624. doi: 10.1128/mbio.01496-24. Epub 2024 Jul 2. mBio. 2024. PMID: 38953635 Free PMC article.
-
Cdk8 and Ssn801 Regulate Oxidative Stress Resistance and Virulence in Cryptococcus neoformans.mBio. 2019 Feb 12;10(1):e02818-18. doi: 10.1128/mBio.02818-18. mBio. 2019. PMID: 30755515 Free PMC article.
-
Inbred Mouse Models in Cryptococcus neoformans Research.J Fungi (Basel). 2024 Jun 17;10(6):426. doi: 10.3390/jof10060426. J Fungi (Basel). 2024. PMID: 38921412 Free PMC article. Review.
References
-
- Andama AO, den Boon S, Meya D, Cattamanchi A, Worodria W, Davis JL, Walter ND, Yoo SD, Kalema N, Haller B, Huang L, International HIV-Associated Opportunistic Pneumonias Study . 2013. Prevalence and outcomes of cryptococcal antigenemia in HIV-seropositive patients hospitalized for suspected tuberculosis in Uganda. J Acquir Immune Defic Syndr 63:189–194. doi:10.1097/QAI.0b013e3182926f95. - DOI - PMC - PubMed
-
- Heitman J, Kozel TR, Kwon-Chung KJ, Perfect JR, Casadevall A. 2010. Cryptococcus: from human pathogen to model yeast. ASM Press, Washington, DC.
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

