Interplay between PTB and miR-1285 at the p53 3'UTR modulates the levels of p53 and its isoform Δ40p53α
- PMID: 28973454
- PMCID: PMC5622325
- DOI: 10.1093/nar/gkx630
Interplay between PTB and miR-1285 at the p53 3'UTR modulates the levels of p53 and its isoform Δ40p53α
Abstract
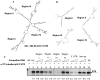
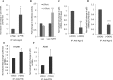
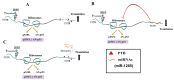
p53 and its translational isoform Δ40p53 are involved in many important cellular functions like cell cycle, cell proliferation, differentiation and metabolism. Expression of both the isoforms can be regulated at different steps. In this study, we explored the role of 3'UTR in regulating the expression of these two translational isoforms. We report that the trans acting factor, Polypyrimidine Tract Binding protein (PTB), also interacts specifically with 3'UTR of p53 mRNA and positively regulates expression of p53 isoforms. Our results suggest that there is interplay between miRNAs and PTB at the 3'UTR under normal and stress conditions like DNA damage. Interestingly, PTB showed some overlapping binding regions in the p53 3'UTR with miR-1285. In fact, knockdown of miR-1285 as well as expression of p53 3'UTR with mutated miR-1285 binding sites resulted in enhanced association of PTB with the 3'UTR, which provides mechanistic insights of this interplay. Taken together, the results provide a plausible molecular basis of how the interplay between miRNAs and the PTB protein at the 3'UTR can play pivotal role in fine tuning the expression of the two p53 isoforms.
© The Author(s) 2017. Published by Oxford University Press on behalf of Nucleic Acids Research.
Figures




Similar articles
-
PTB binds to the 3' untranslated region of the human astrovirus type 8: a possible role in viral replication.PLoS One. 2014 Nov 18;9(11):e113113. doi: 10.1371/journal.pone.0113113. eCollection 2014. PLoS One. 2014. PMID: 25406089 Free PMC article.
-
Delineation of a novel pathway that regulates CD154 (CD40 ligand) expression.Mol Cell Biol. 2003 Jan;23(2):510-25. doi: 10.1128/MCB.23.2.510-525.2003. Mol Cell Biol. 2003. PMID: 12509450 Free PMC article.
-
Polypyrimidine tract binding protein regulates IRES-mediated translation of p53 isoforms.Cell Cycle. 2008 Jul 15;7(14):2189-98. doi: 10.4161/cc.7.14.6271. Epub 2008 May 11. Cell Cycle. 2008. PMID: 18635961
-
The emerging theme of 3'UTR mRNA isoform regulation in reprogramming of cell metabolism.Biochem Soc Trans. 2023 Jun 28;51(3):1111-1119. doi: 10.1042/BST20221128. Biochem Soc Trans. 2023. PMID: 37171086 Free PMC article. Review.
-
Regulation of tumor suppressor p53 at the RNA level.J Mol Med (Berl). 2010 Jul;88(7):645-52. doi: 10.1007/s00109-010-0609-2. Epub 2010 Mar 21. J Mol Med (Berl). 2010. PMID: 20306257 Review.
Cited by
-
Repression of Smad4 by MicroRNA-1285 moderates TGF-β-induced epithelial-mesenchymal transition in proliferative vitreoretinopathy.PLoS One. 2021 Aug 12;16(8):e0254873. doi: 10.1371/journal.pone.0254873. eCollection 2021. PLoS One. 2021. PMID: 34383767 Free PMC article.
-
Re-appraising the evidence for the source, regulation and function of p53-family isoforms.Nucleic Acids Res. 2024 Nov 11;52(20):12112-12129. doi: 10.1093/nar/gkae855. Nucleic Acids Res. 2024. PMID: 39404067 Free PMC article. Review.
-
miR-1285-3p targets TPI1 to regulate the glycolysis metabolism signaling pathway of Tibetan sheep Sertoli cells.PLoS One. 2022 Sep 22;17(9):e0270364. doi: 10.1371/journal.pone.0270364. eCollection 2022. PLoS One. 2022. PMID: 36137140 Free PMC article.
-
The p53 mRNA: an integral part of the cellular stress response.Nucleic Acids Res. 2019 Apr 23;47(7):3257-3271. doi: 10.1093/nar/gkz124. Nucleic Acids Res. 2019. PMID: 30828720 Free PMC article. Review.
-
Conformational stability and dynamics of the cancer-associated isoform Δ133p53β are modulated by p53 peptides and p53-specific DNA.FASEB J. 2019 Mar;33(3):4225-4235. doi: 10.1096/fj.201801973R. Epub 2018 Dec 12. FASEB J. 2019. PMID: 30540922 Free PMC article.
References
-
- Levine A.J. p53, the cellular gatekeeper for growth and division. Cell. 1997; 88:323–331. - PubMed
-
- Candeias M.M., Powell D.J., Roubalova E., Apcher S., Bourougaa K., Vojtesek B., Bruzzoni-Giovanelli H., Fahraeus R.. Expression of p53 and p53/47 are controlled by alternative mechanisms of messenger RNA translation initiation. Oncogene. 2006; 25:6936–6947. - PubMed
-
- Courtois S., Verhaegh G., North S., Luciani M.G., Lassus P., Hibner U., Oren M., Hainaut P.. DeltaN-p53, a natural isoform of P53 lacking the first transactivation domain, counteracts growth suppression by wild-type p53. Oncogene. 2002; 21:6722–6728. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

