The intricate network between the p34 and p44 subunits is central to the activity of the transcription/DNA repair factor TFIIH
- PMID: 28977422
- PMCID: PMC5737387
- DOI: 10.1093/nar/gkx743
The intricate network between the p34 and p44 subunits is central to the activity of the transcription/DNA repair factor TFIIH
Abstract
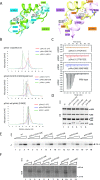
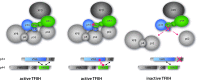
The general transcription factor IIH (TFIIH) is a multi-protein complex and its 10 subunits are engaged in an intricate protein-protein interaction network critical for the regulation of its transcription and DNA repair activities that are so far little understood on a molecular level. In this study, we focused on the p44 and the p34 subunits, which are central for the structural integrity of core-TFIIH. We solved crystal structures of a complex formed by the p34 N-terminal vWA and p44 C-terminal zinc binding domains from Chaetomium thermophilum and from Homo sapiens. Intriguingly, our functional analyses clearly revealed the presence of a second interface located in the C-terminal zinc binding region of p34, which can rescue a disrupted interaction between the p34 vWA and the p44 RING domain. In addition, we demonstrate that the C-terminal zinc binding domain of p34 assumes a central role with respect to the stability and function of TFIIH. Our data reveal a redundant interaction network within core-TFIIH, which may serve to minimize the susceptibility to mutational impairment. This provides first insights why so far no mutations in the p34 or p44 TFIIH-core subunits have been identified that would lead to the hallmark nucleotide excision repair syndromes xeroderma pigmentosum or trichothiodystrophy.
© The Author(s) 2017. Published by Oxford University Press on behalf of Nucleic Acids Research.
Figures



Similar articles
-
The structure of the TFIIH p34 subunit reveals a von Willebrand factor A like fold.PLoS One. 2014 Jul 11;9(7):e102389. doi: 10.1371/journal.pone.0102389. eCollection 2014. PLoS One. 2014. PMID: 25013903 Free PMC article.
-
The cryo-electron microscopy structure of human transcription factor IIH.Nature. 2017 Sep 21;549(7672):414-417. doi: 10.1038/nature23903. Epub 2017 Sep 13. Nature. 2017. PMID: 28902838 Free PMC article.
-
How to limit the speed of a motor: the intricate regulation of the XPB ATPase and translocase in TFIIH.Nucleic Acids Res. 2020 Dec 2;48(21):12282-12296. doi: 10.1093/nar/gkaa911. Nucleic Acids Res. 2020. PMID: 33196848 Free PMC article.
-
Nucleotide Excision Repair and Transcriptional Regulation: TFIIH and Beyond.Annu Rev Biochem. 2016 Jun 2;85:265-90. doi: 10.1146/annurev-biochem-060815-014857. Annu Rev Biochem. 2016. PMID: 27294439 Review.
-
A history of TFIIH: two decades of molecular biology on a pivotal transcription/repair factor.DNA Repair (Amst). 2011 Jul 15;10(7):714-21. doi: 10.1016/j.dnarep.2011.04.021. Epub 2011 May 17. DNA Repair (Amst). 2011. PMID: 21592869 Review.
Cited by
-
HR-Bac, a toolbox based on homologous recombination for expression, screening and production of multiprotein complexes using the baculovirus expression system.Sci Rep. 2022 Feb 7;12(1):2030. doi: 10.1038/s41598-021-04715-5. Sci Rep. 2022. PMID: 35132103 Free PMC article.
-
The complete structure of the human TFIIH core complex.Elife. 2019 Mar 12;8:e44771. doi: 10.7554/eLife.44771. Elife. 2019. PMID: 30860024 Free PMC article.
-
The essential and multifunctional TFIIH complex.Protein Sci. 2018 Jun;27(6):1018-1037. doi: 10.1002/pro.3424. Epub 2018 Apr 27. Protein Sci. 2018. PMID: 29664212 Free PMC article. Review.
-
Transcription preinitiation complex structure and dynamics provide insight into genetic diseases.Nat Struct Mol Biol. 2019 Jun;26(6):397-406. doi: 10.1038/s41594-019-0220-3. Epub 2019 May 20. Nat Struct Mol Biol. 2019. PMID: 31110295 Free PMC article.
-
Nucleotide Excision Repair: Insights into Canonical and Emerging Functions of the Transcription/DNA Repair Factor TFIIH.Genes (Basel). 2025 Feb 19;16(2):231. doi: 10.3390/genes16020231. Genes (Basel). 2025. PMID: 40004560 Free PMC article. Review.
References
-
- Compe E., Egly J.M.. Nucleotide excision repair and transcriptional regulation: TFIIH and beyond. Annu. Rev. Biochem. 2016; 85:265–290. - PubMed
-
- Cleaver J.E. Splitting hairs–discovery of a new DNA repair and transcription factor for the human disease trichothiodystrophy. DNA Repair (Amst). 2005; 4:285–287. - PubMed
-
- Lehmann A.R., Norris P.G.. DNA repair and cancer: speculations based on studies with xeroderma pigmentosum, Cockayne's syndrome and trichothiodystrophy. Carcinogenesis. 1989; 10:1353–1356. - PubMed
-
- Yoon H., Miller S.P., Pabich E.K., Donahue T.F.. SSL1, a suppressor of a HIS4 5′-UTR stem-loop mutation, is essential for translation initiation and affects UV resistance in yeast. Genes Dev. 1992; 6:2463–2477. - PubMed
-
- Gulyas K.D., Donahue T.F.. SSL2, a suppressor of a stem-loop mutation in the HIS4 leader encodes the yeast homolog of human ERCC-3. Cell. 1992; 69:1031–1042. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

