Epithelial-Myeloid cell crosstalk regulates acinar cell plasticity and pancreatic remodeling in mice
- PMID: 28980940
- PMCID: PMC5690281
- DOI: 10.7554/eLife.27388
Epithelial-Myeloid cell crosstalk regulates acinar cell plasticity and pancreatic remodeling in mice
Abstract
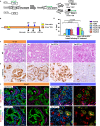
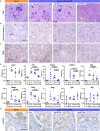
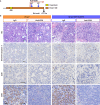
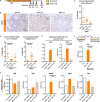
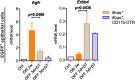
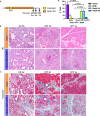
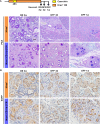
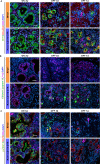
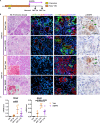
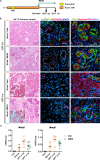
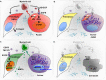
Dedifferentiation of acini to duct-like cells occurs during the physiologic damage response in the pancreas, but this process can be co-opted by oncogenic Kras to drive carcinogenesis. Myeloid cells infiltrate the pancreas during the onset of pancreatic cancer, and promote carcinogenesis. Here, we show that the function of infiltrating myeloid cells is regulated by oncogenic Kras expressed in epithelial cells. In the presence of oncogenic Kras, myeloid cells promote acinar dedifferentiation and carcinogenesis. Upon inactivation of oncogenic Kras, myeloid cells promote re-differentiation of acinar cells, remodeling of the fibrotic stroma and tissue repair. Intriguingly, both aspects of myeloid cell activity depend, at least in part, on activation of EGFR/MAPK signaling, with different subsets of ligands and receptors in different target cells promoting carcinogenesis or repair, respectively. Thus, the cross-talk between epithelial cells and infiltrating myeloid cells determines the balance between tissue repair and carcinogenesis in the pancreas.
Keywords: EGF; MAPK; cancer biology; mouse; myeloid cell; pancreatic cancer; signaling pathways; tissue remodeling.
Conflict of interest statement
No competing interests declared.
Figures

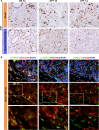
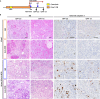
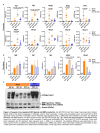
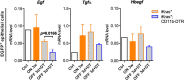
References
-
- Ardito CM, Grüner BM, Takeuchi KK, Lubeseder-Martellato C, Teichmann N, Mazur PK, Delgiorno KE, Carpenter ES, Halbrook CJ, Hall JC, Pal D, Briel T, Herner A, Trajkovic-Arsic M, Sipos B, Liou GY, Storz P, Murray NR, Threadgill DW, Sibilia M, Washington MK, Wilson CL, Schmid RM, Raines EW, Crawford HC, Siveke JT. EGF receptor is required for KRAS-induced pancreatic tumorigenesis. Cancer Cell. 2012;22:304–317. doi: 10.1016/j.ccr.2012.07.024. - DOI - PMC - PubMed
-
- Bailey P, Chang DK, Nones K, Johns AL, Patch AM, Gingras MC, Miller DK, Christ AN, Bruxner TJ, Quinn MC, Nourse C, Murtaugh LC, Harliwong I, Idrisoglu S, Manning S, Nourbakhsh E, Wani S, Fink L, Holmes O, Chin V, Anderson MJ, Kazakoff S, Leonard C, Newell F, Waddell N, Wood S, Xu Q, Wilson PJ, Cloonan N, Kassahn KS, Taylor D, Quek K, Robertson A, Pantano L, Mincarelli L, Sanchez LN, Evers L, Wu J, Pinese M, Cowley MJ, Jones MD, Colvin EK, Nagrial AM, Humphrey ES, Chantrill LA, Mawson A, Humphris J, Chou A, Pajic M, Scarlett CJ, Pinho AV, Giry-Laterriere M, Rooman I, Samra JS, Kench JG, Lovell JA, Merrett ND, Toon CW, Epari K, Nguyen NQ, Barbour A, Zeps N, Moran-Jones K, Jamieson NB, Graham JS, Duthie F, Oien K, Hair J, Grützmann R, Maitra A, Iacobuzio-Donahue CA, Wolfgang CL, Morgan RA, Lawlor RT, Corbo V, Bassi C, Rusev B, Capelli P, Salvia R, Tortora G, Mukhopadhyay D, Petersen GM, Munzy DM, Fisher WE, Karim SA, Eshleman JR, Hruban RH, Pilarsky C, Morton JP, Sansom OJ, Scarpa A, Musgrove EA, Bailey UM, Hofmann O, Sutherland RL, Wheeler DA, Gill AJ, Gibbs RA, Pearson JV, Waddell N, Biankin AV, Grimmond SM, Australian Pancreatic Cancer Genome Initiative Genomic analyses identify molecular subtypes of pancreatic cancer. Nature. 2016;531:47–52. doi: 10.1038/nature16965. - DOI - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous

