Integrated microarray analysis provided novel insights to the pathogenesis of glaucoma
- PMID: 28990066
- PMCID: PMC5779953
- DOI: 10.3892/mmr.2017.7711
Integrated microarray analysis provided novel insights to the pathogenesis of glaucoma
Abstract
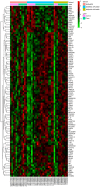
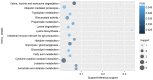
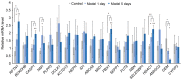
Glaucoma is characterized as a visual field defect, which is the second most common cause of blindness. The present study performed an integrated analysis of microarray studies of glaucoma derived from Gene Expression Omnibus (GEO). Following the identification of the differentially expressed genes (DEGs) in glaucoma compared with normal control (NC) tissues, the functional annotation, glaucoma‑specific protein‑protein interaction (PPI) network and transcriptional regulatory network constructions were performed. The acute intraocular pressure (IOP) elevation rat models were established and reverse transcription‑quantitative polymerase chain reaction (RT‑qPCR) was performed for DEGs expression confirmation. Three datasets were downloaded from GEO. A total of 97 DEGs, 82 upregulated and 15 downregulated were identified in glaucoma compared with NC groups with false discovery rate <0.05. Response to virus and immune response were two significantly enriched GO terms in glaucoma. Valine, leucine and isoleucine degradation was a significantly enriched pathway of DEGs in glaucoma. According to the PPI network, HDAC1, HBN, UBR4 and PDK1 were hub proteins in glaucoma. FOXD3, HNF‑4 and AP‑1 were the three transcription factors (TFs) derived from top 10 TFs which covered the majority of downstream DEGs in glaucoma. Based on the RT‑qPCR results, the expression levels of 3 DEGs, raftlin, lipid raft linker 1 (RFTN1), PBX homeobox 1 (PBX1), HDAC1 were significantly upregulated and the expression of GEM was significantly downregulated in acute IOP elevation rat model at the first and fifth day. These four DEGs had the same expression pattern with our integrated analysis. Therefore, the current study concluded that 6 DEGs, including HEPH, SELENBP1, RFTN1, ID1, HDAC‑1 and PBX1 and three TFs, including FOXD3, HNF‑4 and AP‑1 may be involved with the pathogenesis of glaucoma. The findings of the current study may improve diagnosis and drug design for glaucoma.
Figures





References
-
- Moreno MC, Sande P, Marcos HA, de Zavalía N, Sarmiento MI Keller, Rosenstein RE. Effect of glaucoma on the retinal glutamate/glutamine cycle activity. FASEB J. 2005;19:1161–1162. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials
Miscellaneous

