An NAM Domain Gene, GhNAC79, Improves Resistance to Drought Stress in Upland Cotton
- PMID: 28993786
- PMCID: PMC5622203
- DOI: 10.3389/fpls.2017.01657
An NAM Domain Gene, GhNAC79, Improves Resistance to Drought Stress in Upland Cotton
Abstract
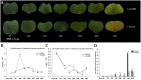
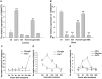
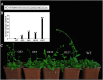
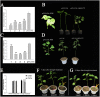
Plant-specific NAC proteins comprise one of the largest transcription factor families in plants and play important roles in plant development and the stress response. Gossypium hirsutum L. is a major source of fiber, but its growth and productivity are limited by many biotic and abiotic stresses. In this study, the NAC domain gene GhNAC79 was functionally characterized in detail, and according to information about the cotton genome sequences, it was located on scaffold42.1, containing three exons and two introns. Promoter analysis indicated that the GhNAC79 promoter contained both basic and stress-related elements, and it was especially expressed in the cotyledon of Arabidopsis. A transactivation assay in yeast demonstrated that GhNAC79 was a transcription activator, and its activation domain was located at its C-terminus. The results of qRT-PCR proved that GhNAC79 was preferentially expressed at later stages of cotyledon and fiber development, and it showed high sensitivity to ethylene and meJA treatments. Overexpression of GhNAC79 resulted in an early flowering phenotype in Arabidopsis, and it also improved drought tolerance in both Arabidopsis and cotton. Furthermore, VIGS-induced silencing of GhNAC79 in cotton led to a drought-sensitive phenotype. In summary, GhNAC79 positively regulates drought stress, and it also responds to ethylene and meJA treatments, making it a candidate gene for stress studies in cotton.
Keywords: GhNAC79; cotton; development; drought; stress.
Figures



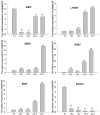
References
-
- Bartels D., Sunkar R. (2005). Drought and salt tolerance in plants. Crit. Rev. Plant Sci. 24 23–58. 10.1080/07352680590910410 - DOI
LinkOut - more resources
Full Text Sources
Other Literature Sources

