A recombinant lentiviral PDGF-driven mouse model of proneural glioblastoma
- PMID: 29016807
- PMCID: PMC5817944
- DOI: 10.1093/neuonc/nox129
A recombinant lentiviral PDGF-driven mouse model of proneural glioblastoma
Abstract
Background: Mouse models of glioblastoma (GBM), the most aggressive primary brain tumor, are critical for understanding GBM pathology and can contribute to the preclinical evaluation of therapeutic agents. Platelet-derived growth factor (PDGF) signaling has been implicated in the development and pathogenesis of GBM, specifically the proneural subtype. Although multiple mouse models of PDGF-driven glioma have been described, they require transgenic mice engineered to activate PDGF signaling and/or impair tumor suppressor genes and typically represent lower-grade glioma.
Methods: We designed recombinant lentiviruses expressing both PDGFB and a short hairpin RNA targeting Cdkn2a to induce gliomagenesis following stereotactic injection into the dentate gyrus of adult immunocompetent mice. We engineered these viruses to coexpress CreERT2 with PDGFB, allowing for deletion of floxed genes specifically in transduced cells, and designed another version of this recombinant lentivirus in which enhanced green fluorescent protein was coexpressed with PDGFB and CreERT2 to visualize transduced cells.
Results: The dentate gyrus of injected mice showed hypercellularity one week post-injection and subsequently developed bona fide tumors with the pathologic hallmarks of GBM leading to a median survival of 77 days post-injection. Transcriptomic analysis of these tumors revealed a proneural gene expression signature.
Conclusion: Informed by the genetic alterations observed in human GBM, we engineered a novel mouse model of proneural GBM. While reflecting many of the advantages of transgenic mice, this model allows for the facile in vivo testing of gene function in tumor cells and makes possible the rapid production of large numbers of immunocompetent tumor-bearing mice for preclinical testing of therapeutics.
© The Author(s) 2017. Published by Oxford University Press on behalf of the Society for Neuro-Oncology. All rights reserved. For permissions, please e-mail: journals.permissions@oup.com
Figures
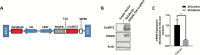





Comment in
-
A new practical and versatile mouse model of proneural glioblastoma.Neuro Oncol. 2018 Feb 19;20(3):299-301. doi: 10.1093/neuonc/nox233. Neuro Oncol. 2018. PMID: 29432618 Free PMC article. No abstract available.
Similar articles
-
The HIF1α-PDGFD-PDGFRα axis controls glioblastoma growth at normoxia/mild-hypoxia and confers sensitivity to targeted therapy by echinomycin.J Exp Clin Cancer Res. 2021 Sep 1;40(1):278. doi: 10.1186/s13046-021-02082-7. J Exp Clin Cancer Res. 2021. PMID: 34470658 Free PMC article.
-
MicroRNA-29a inhibits glioblastoma stem cells and tumor growth by regulating the PDGF pathway.J Neurooncol. 2019 Oct;145(1):23-34. doi: 10.1007/s11060-019-03275-z. Epub 2019 Sep 3. J Neurooncol. 2019. PMID: 31482267 Free PMC article.
-
PDGF Engages an E2F-USP1 Signaling Pathway to Support ID2-Mediated Survival of Proneural Glioma Cells.Cancer Res. 2016 May 15;76(10):2964-76. doi: 10.1158/0008-5472.CAN-15-2157. Epub 2016 Mar 7. Cancer Res. 2016. PMID: 26951930
-
Platelet-derived growth factor in glioblastoma-driver or biomarker?Ups J Med Sci. 2014 Nov;119(4):298-305. doi: 10.3109/03009734.2014.970304. Epub 2014 Oct 24. Ups J Med Sci. 2014. PMID: 25342206 Free PMC article. Review.
-
Molecular and Genomic Alterations in Glioblastoma Multiforme.Am J Pathol. 2015 Jul;185(7):1820-33. doi: 10.1016/j.ajpath.2015.02.023. Epub 2015 May 11. Am J Pathol. 2015. PMID: 25976245 Review.
Cited by
-
Comprehensive understanding of glioblastoma molecular phenotypes: classification, characteristics, and transition.Cancer Biol Med. 2024 May 6;21(5):363-81. doi: 10.20892/j.issn.2095-3941.2023.0510. Cancer Biol Med. 2024. PMID: 38712813 Free PMC article. Review.
-
Expert opinion on translational research for advanced glioblastoma treatment.Cancer Biol Med. 2023 Apr 25;20(5):344-52. doi: 10.20892/j.issn.2095-3941.2023.0012. Cancer Biol Med. 2023. PMID: 37092846 Free PMC article. Review.
-
Evolving cell states and oncogenic drivers during the progression of IDH-mutant gliomas.Nat Cancer. 2025 Jan;6(1):145-157. doi: 10.1038/s43018-024-00865-3. Epub 2024 Nov 21. Nat Cancer. 2025. PMID: 39572850
-
EPHA2 mediates PDGFA activity and functions together with PDGFRA as prognostic marker and therapeutic target in glioblastoma.Signal Transduct Target Ther. 2022 Feb 2;7(1):33. doi: 10.1038/s41392-021-00855-2. Signal Transduct Target Ther. 2022. PMID: 35105853 Free PMC article.
-
The HIF1α-PDGFD-PDGFRα axis controls glioblastoma growth at normoxia/mild-hypoxia and confers sensitivity to targeted therapy by echinomycin.J Exp Clin Cancer Res. 2021 Sep 1;40(1):278. doi: 10.1186/s13046-021-02082-7. J Exp Clin Cancer Res. 2021. PMID: 34470658 Free PMC article.
References
-
- Omuro A, DeAngelis LM. Glioblastoma and other malignant gliomas: a clinical review. JAMA. 2013;310(17):1842–1850. - PubMed
-
- Louis DN, Perry A, Reifenberger G et al. . The 2016 World Health Organization classification of tumors of the central nervous system: a summary. Acta Neuropathol. 2016;131(6):803–820. - PubMed
-
- Aldape K, Zadeh G, Mansouri S, Reifenberger G, von Deimling A. Glioblastoma: pathology, molecular mechanisms and markers. Acta Neuropathol. 2015;129(6):829–848. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Miscellaneous

