Identification of 76 novel B1 metallo-β-lactamases through large-scale screening of genomic and metagenomic data
- PMID: 29020980
- PMCID: PMC5637372
- DOI: 10.1186/s40168-017-0353-8
Identification of 76 novel B1 metallo-β-lactamases through large-scale screening of genomic and metagenomic data
Abstract
Background: Metallo-β-lactamases are bacterial enzymes that provide resistance to carbapenems, the most potent class of antibiotics. These enzymes are commonly encoded on mobile genetic elements, which, together with their broad substrate spectrum and lack of clinically useful inhibitors, make them a particularly problematic class of antibiotic resistance determinants. We hypothesized that there is a large and unexplored reservoir of unknown metallo-β-lactamases, some of which may spread to pathogens, thereby threatening public health. The aim of this study was to identify novel metallo-β-lactamases of class B1, the most clinically important subclass of these enzymes.
Results: Based on a new computational method using an optimized hidden Markov model, we analyzed over 10,000 bacterial genomes and plasmids together with more than 5 terabases of metagenomic data to identify novel metallo-β-lactamase genes. In total, 76 novel genes were predicted, forming 59 previously undescribed metallo-β-lactamase gene families. The ability to hydrolyze imipenem in an Escherichia coli host was experimentally confirmed for 18 of the 21 tested genes. Two of the novel B1 metallo-β-lactamase genes contained atypical zinc-binding motifs in their active sites, which were previously undescribed for metallo-β-lactamases. Phylogenetic analysis showed that B1 metallo-β-lactamases could be divided into five major groups based on their evolutionary origin. Our results also show that, except for one, all of the previously characterized mobile B1 β-lactamases are likely to have originated from chromosomal genes present in Shewanella spp. and other Proteobacterial species.
Conclusions: This study more than doubles the number of known B1 metallo-β-lactamases. The findings have further elucidated the diversity and evolutionary history of this important class of antibiotic resistance genes and prepare us for some of the challenges that may be faced in clinics in the future.
Keywords: Carbapenemases; Hidden Markov model; Resistome; Shotgun metagenomics; β-lactam resistance.
Conflict of interest statement
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Figures
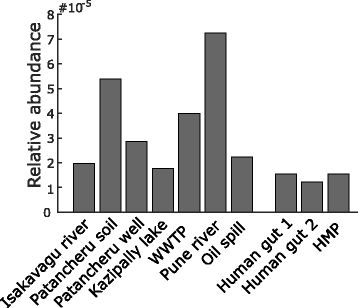
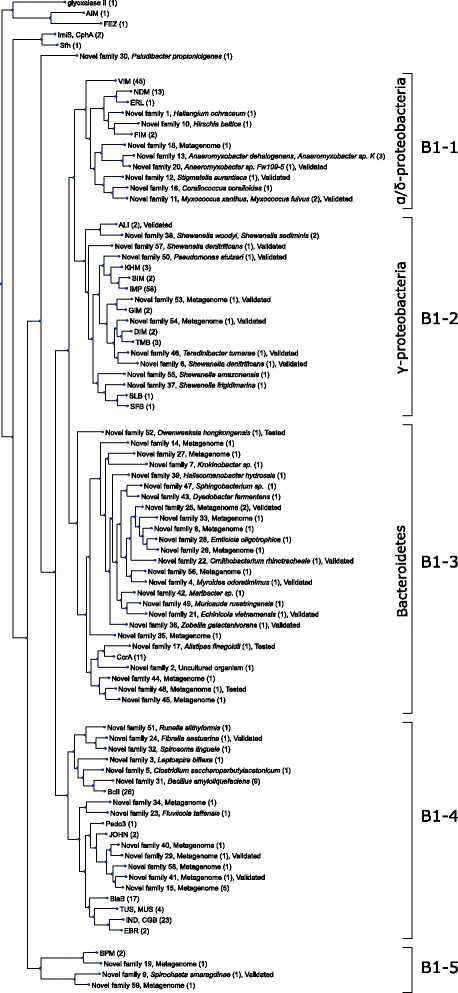
Similar articles
-
Sewage effluent from an Indian hospital harbors novel carbapenemases and integron-borne antibiotic resistance genes.Microbiome. 2019 Jun 27;7(1):97. doi: 10.1186/s40168-019-0710-x. Microbiome. 2019. PMID: 31248462 Free PMC article.
-
Metallo-beta-lactamases (classification, activity, genetic organization, structure, zinc coordination) and their superfamily.Biochem Pharmacol. 2007 Dec 15;74(12):1686-701. doi: 10.1016/j.bcp.2007.05.021. Epub 2007 Jun 2. Biochem Pharmacol. 2007. PMID: 17597585 Review.
-
An updated phylogeny of the metallo-β-lactamases.J Antimicrob Chemother. 2021 Jan 1;76(1):117-123. doi: 10.1093/jac/dkaa392. J Antimicrob Chemother. 2021. PMID: 33005957
-
What's new in antibiotic resistance? Focus on beta-lactamases.Drug Resist Updat. 2006 Jun;9(3):142-56. doi: 10.1016/j.drup.2006.05.005. Epub 2006 Aug 8. Drug Resist Updat. 2006. PMID: 16899402 Review.
-
The Reaction Mechanism of Metallo-β-Lactamases Is Tuned by the Conformation of an Active-Site Mobile Loop.Antimicrob Agents Chemother. 2018 Dec 21;63(1):e01754-18. doi: 10.1128/AAC.01754-18. Print 2019 Jan. Antimicrob Agents Chemother. 2018. PMID: 30348667 Free PMC article.
Cited by
-
Systematic Identification and Classification of β-Lactamases Based on Sequence Similarity Criteria: β-Lactamase Annotation.Evol Bioinform Online. 2018 Sep 10;14:1176934318797351. doi: 10.1177/1176934318797351. eCollection 2018. Evol Bioinform Online. 2018. PMID: 30210232 Free PMC article.
-
Molecular and Functional Characterization of a Novel Plasmid-Borne bla NDM-Like Gene, bla AFM-1, in a Clinical Strain of Aeromonas hydrophila.Infect Drug Resist. 2021 Apr 22;14:1613-1622. doi: 10.2147/IDR.S297419. eCollection 2021. Infect Drug Resist. 2021. PMID: 33911885 Free PMC article.
-
Metagenomic Screening for Lipolytic Genes Reveals an Ecology-Clustered Distribution Pattern.Front Microbiol. 2022 Jun 10;13:851969. doi: 10.3389/fmicb.2022.851969. eCollection 2022. Front Microbiol. 2022. PMID: 35756004 Free PMC article.
-
Characterization of blaAFM-1-positive carbapenem-resistant strains isolated in Guangzhou, China.Ann Clin Microbiol Antimicrob. 2023 May 17;22(1):40. doi: 10.1186/s12941-023-00592-0. Ann Clin Microbiol Antimicrob. 2023. PMID: 37198688 Free PMC article.
-
Novel Antibiotic Resistance Genes Identified by Functional Gene Library Screening in Stenotrophomonas maltophilia and Chryseobacterium spp. Bacteria of Soil Origin.Int J Mol Sci. 2023 Mar 23;24(7):6037. doi: 10.3390/ijms24076037. Int J Mol Sci. 2023. PMID: 37047008 Free PMC article.
References
-
- Fast W, Sutton LD. Metallo-beta-lactamase: inhibitors and reporter substrates. Biochim Biophys Acta. 1834;2013:1648–1659. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

