Amyloidogenic motifs revealed by n-gram analysis
- PMID: 29021608
- PMCID: PMC5636826
- DOI: 10.1038/s41598-017-13210-9
Amyloidogenic motifs revealed by n-gram analysis
Abstract
Amyloids are proteins associated with several clinical disorders, including Alzheimer's, and Creutzfeldt-Jakob's. Despite their diversity, all amyloid proteins can undergo aggregation initiated by short segments called hot spots. To find the patterns defining the hot spots, we trained predictors of amyloidogenicity, using n-grams and random forest classifiers. Since the amyloidogenicity may not depend on the exact sequence of amino acids but on their more general properties, we tested 524,284 reduced amino acid alphabets of different lengths (three to six letters) to find the alphabet providing the best performance in cross-validation. The predictor based on this alphabet, called AmyloGram, was benchmarked against the most popular tools for the detection of amyloid peptides using an external data set and obtained the highest values of performance measures (AUC: 0.90, MCC: 0.63). Our results showed sequential patterns in the amyloids which are strongly correlated with hydrophobicity, a tendency to form β-sheets, and lower flexibility of amino acid residues. Among the most informative n-grams of AmyloGram we identified 15 that were previously confirmed experimentally. AmyloGram is available as the web-server: http://smorfland.uni.wroc.pl/shiny/AmyloGram/ and as the R package AmyloGram. R scripts and data used to produce the results of this manuscript are available at http://github.com/michbur/AmyloGramAnalysis .
Conflict of interest statement
The authors declare that they have no competing interests.
Figures
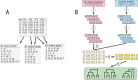



References
-
- Chaturvedi, S. K., Siddiqi, M. K., Alam, P. & Khan, R. H. Protein misfolding and aggregation: Mechanism, factors and detection. Process. Biochem. 51(9), 1183–1192 (2016).
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

