Dynamic DNA binding, junction recognition and G4 melting activity underlie the telomeric and genome-wide roles of human CST
- PMID: 29040642
- PMCID: PMC5716219
- DOI: 10.1093/nar/gkx878
Dynamic DNA binding, junction recognition and G4 melting activity underlie the telomeric and genome-wide roles of human CST
Abstract
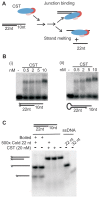
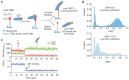
Human CST (CTC1-STN1-TEN1) is a ssDNA-binding complex that helps resolve replication problems both at telomeres and genome-wide. CST resembles Replication Protein A (RPA) in that the two complexes harbor comparable arrays of OB-folds and have structurally similar small subunits. However, the overall architecture and functions of CST and RPA are distinct. Currently, the mechanism underlying CST action at diverse replication issues remains unclear. To clarify CST mechanism, we examined the capacity of CST to bind and resolve DNA structures found at sites of CST activity. We show that CST binds preferentially to ss-dsDNA junctions, an activity that can explain the incremental nature of telomeric C-strand synthesis following telomerase action. We also show that CST unfolds G-quadruplex structures, thus providing a mechanism for CST to facilitate replication through telomeres and other GC-rich regions. Finally, smFRET analysis indicates that CST binding to ssDNA is dynamic with CST complexes undergoing concentration-dependent self-displacement. These findings support an RPA-based model where dissociation and re-association of individual OB-folds allow CST to mediate loading and unloading of partner proteins to facilitate various aspects of telomere replication and genome-wide resolution of replication stress.
© The Author(s) 2017. Published by Oxford University Press on behalf of Nucleic Acids Research.
Figures





References
-
- Miyake Y., Nakamura M., Nabetani A., Shimamura S., Tamura M., Yonehara S., Saito M., Ishikawa F.. RPA-like mammalian CTC1–STN1–TEN1 complex binds to single-stranded DNA and protects telomeres independently of the Pot1 pathway. Mol. Cell. 2009; 36:193–206. - PubMed
-
- Chen L.Y., Redon S., Lingner J.. The human CST complex is a terminator of telomerase activity. Nature. 2012; 488:540–544. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

