Calcium-driven DNA synthesis by a high-fidelity DNA polymerase
- PMID: 29040737
- PMCID: PMC5716173
- DOI: 10.1093/nar/gkx927
Calcium-driven DNA synthesis by a high-fidelity DNA polymerase
Abstract
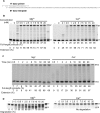
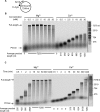
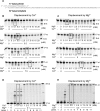
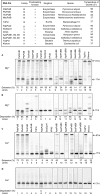
Divalent metal ions, usually Mg2+, are required for both DNA synthesis and proofreading functions by DNA polymerases (DNA Pol). Although used as a non-reactive cofactor substitute for binding and crystallographic studies, Ca2+ supports DNA polymerization by only one DNA Pol, Dpo4. Here, we explore whether Ca2+-driven catalysis might apply to high-fidelity (HiFi) family B DNA Pols. The consequences of replacing Mg2+ by Ca2+ on base pairing at the polymerase active site as well as the editing of terminal nucleotides at the exonuclease active site of the archaeal Pyrococcus abyssi DNA Pol (PabPolB) are characterized and compared to other (families B, A, Y, X, D) DNA Pols. Based on primer extension assays, steady-state kinetics and ion-chased experiments, we demonstrate that Ca2+ (and other metal ions) activates DNA synthesis by PabPolB. While showing a slower rate of phosphodiester bond formation, nucleotide selectivity is improved over that of Mg2+. Further mechanistic studies show that the affinities for primer/template are higher in the presence of Ca2+ and reinforced by a correct incoming nucleotide. Conversely, no exonuclease degradation of the terminal nucleotides occurs with Ca2+. Evolutionary and mechanistic insights among DNA Pols are thus discussed.
© The Author(s) 2017. Published by Oxford University Press on behalf of Nucleic Acids Research.
Figures


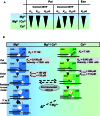
Similar articles
-
The hyperthermophilic euryarchaeota Pyrococcus abyssi likely requires the two DNA polymerases D and B for DNA replication.J Mol Biol. 2005 Jul 1;350(1):53-64. doi: 10.1016/j.jmb.2005.04.042. J Mol Biol. 2005. PMID: 15922358
-
Molecular recognition of canonical and deaminated bases by P. abyssi family B DNA polymerase.J Mol Biol. 2012 Oct 26;423(3):315-36. doi: 10.1016/j.jmb.2012.07.025. Epub 2012 Aug 16. J Mol Biol. 2012. PMID: 22902479
-
Intrinsic properties of the two replicative DNA polymerases of Pyrococcus abyssi in replicating abasic sites: possible role in DNA damage tolerance?Mol Microbiol. 2008 Nov;70(3):746-61. doi: 10.1111/j.1365-2958.2008.06446.x. Epub 2008 Sep 24. Mol Microbiol. 2008. PMID: 18826407
-
Different Divalent Cations Alter the Kinetics and Fidelity of DNA Polymerases.J Biol Chem. 2016 Sep 30;291(40):20869-20875. doi: 10.1074/jbc.R116.742494. Epub 2016 Jul 26. J Biol Chem. 2016. PMID: 27462081 Free PMC article. Review.
-
Conformational coupling in DNA polymerase fidelity.Annu Rev Biochem. 1993;62:685-713. doi: 10.1146/annurev.bi.62.070193.003345. Annu Rev Biochem. 1993. PMID: 7688945 Review.
Cited by
-
Bst polymerase - a humble relative of Taq polymerase.Comput Struct Biotechnol J. 2023 Sep 12;21:4519-4535. doi: 10.1016/j.csbj.2023.09.008. eCollection 2023. Comput Struct Biotechnol J. 2023. PMID: 37767105 Free PMC article. Review.
-
Mechanistic Insights into the Phase Separation Behavior and Pathway-Directed Information Exchange in all-DNA Droplets.Angew Chem Int Ed Engl. 2022 Nov 7;61(45):e202208951. doi: 10.1002/anie.202208951. Epub 2022 Oct 7. Angew Chem Int Ed Engl. 2022. PMID: 36112754 Free PMC article.
-
Liquid drop of DNA libraries reveals total genome information.Proc Natl Acad Sci U S A. 2020 Nov 3;117(44):27300-27306. doi: 10.1073/pnas.2017138117. Epub 2020 Oct 21. Proc Natl Acad Sci U S A. 2020. PMID: 33087570 Free PMC article.
-
Direct Enzyme Engineering of B Family DNA Polymerases for Biotechnological Approaches.Bioengineering (Basel). 2023 Sep 30;10(10):1150. doi: 10.3390/bioengineering10101150. Bioengineering (Basel). 2023. PMID: 37892880 Free PMC article. Review.
-
Synthesis of phosphoramidate-linked DNA by a modified DNA polymerase.Proc Natl Acad Sci U S A. 2020 Mar 31;117(13):7276-7283. doi: 10.1073/pnas.1922400117. Epub 2020 Mar 18. Proc Natl Acad Sci U S A. 2020. PMID: 32188786 Free PMC article.
References
-
- Bebenek K., Kunkel T.A.. Functions of DNA polymerases. Adv. Protein Chem. 2004; 69:137–165. - PubMed
-
- Ishino Y., Ishino S.. DNA replication in Archaea, the third domain of life. Tech, Book series: The Mechanisms of DNA Replication. 2013; Croatia: In Tech Open (E-Books) http://www.intechopen.com/articles/show/title/dna-replication-in-archaea....
-
- Kool E.T. Active site tightness and substrate fit in DNA replication. Annu. Rev. Biochem. 2002; 71:191–219. - PubMed
-
- Rothwell P.J., Waksman G.. Structure and mechanism of DNA polymerases. Adv. Protein Chem. 2005; 71:401–440. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

