The hypothetical protein P47 of Clostridium botulinum E1 strain Beluga has a structural topology similar to bactericidal/permeability-increasing protein
- PMID: 29042313
- PMCID: PMC5902665
- DOI: 10.1016/j.toxicon.2017.10.012
The hypothetical protein P47 of Clostridium botulinum E1 strain Beluga has a structural topology similar to bactericidal/permeability-increasing protein
Abstract
Botulinum neurotoxins (BoNTs) are causative agents of the life-threatening disease botulism. They are naturally produced by species of the bacteria Clostridium botulinum as stable and non-covalent complexes, in which the BoNT molecule is assembled with several auxiliary non-toxic proteins. Some BoNT serotypes, represented by the well-studied BoNT serotype A (BoNT/A), are produced by Clostridium strains that carry the ha gene cluster, which encodes four neurotoxin-associated proteins (NTNHA, HA17, HA33, and HA70) that play an important role to deliver and protect BoNTs in the gastrointestinal tract during oral intoxication. In contrast, BoNT/E- and BoNT/F-producing strains carry a distinct gene cluster that encodes five proteins (NTNHA, P47, OrfX1, OrfX2, and OrfX3, termed the orfX cluster). The structures and functions of these proteins remain largely unknown. Here, we report the crystal structure of P47 resolved at 2.8 Å resolution. Surprisingly, P47 displays a structural topology that is similar to bactericidal/permeability-increasing (BPI) like proteins, which were previously identified only in eukaryotes. The similarity of a hydrophobic cleft of P47 with the phospholipid-binding groove of BPI suggests that P47 might be involved in lipid association to exert its function. Consistently, P47 associates and induces aggregation of asolectin-containing liposomes in a protein- and lipid-concentration dependent manner. These findings laid the foundation for future structural and functional studies of the potential roles of P47 and OrfX proteins in facilitating oral intoxication of BoNTs.
Keywords: Bactericidal/permeability-increasing protein; Botulinum neurotoxin; Crystal structure; Lipid binding; P47; Toxin complex; orfX gene cluster.
Copyright © 2017 Elsevier Ltd. All rights reserved.
Figures
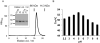

Similar articles
-
Looking for the X Factor in Bacterial Pathogenesis: Association of orfX-p47 Gene Clusters with Toxin Genes in Clostridial and Non-Clostridial Bacterial Species.Toxins (Basel). 2019 Dec 31;12(1):19. doi: 10.3390/toxins12010019. Toxins (Basel). 2019. PMID: 31906154 Free PMC article.
-
Crystal structures of OrfX1, OrfX2 and the OrfX1-OrfX3 complex from the orfX gene cluster of botulinum neurotoxin E1.FEBS Lett. 2023 Feb;597(4):524-537. doi: 10.1002/1873-3468.14576. Epub 2023 Jan 27. FEBS Lett. 2023. PMID: 36653893 Free PMC article.
-
Crystal structures of OrfX2 and P47 from a Botulinum neurotoxin OrfX-type gene cluster.FEBS Lett. 2017 Nov;591(22):3781-3792. doi: 10.1002/1873-3468.12889. Epub 2017 Nov 5. FEBS Lett. 2017. PMID: 29067689
-
[Clostridium botulinum and botulinum neurotoxin].Brain Nerve. 2011 Jul;63(7):755-61. Brain Nerve. 2011. PMID: 21747146 Review. Japanese.
-
Assembly and function of the botulinum neurotoxin progenitor complex.Curr Top Microbiol Immunol. 2013;364:21-44. doi: 10.1007/978-3-642-33570-9_2. Curr Top Microbiol Immunol. 2013. PMID: 23239347 Free PMC article. Review.
Cited by
-
Looking for the X Factor in Bacterial Pathogenesis: Association of orfX-p47 Gene Clusters with Toxin Genes in Clostridial and Non-Clostridial Bacterial Species.Toxins (Basel). 2019 Dec 31;12(1):19. doi: 10.3390/toxins12010019. Toxins (Basel). 2019. PMID: 31906154 Free PMC article.
-
Revisiting the Evolution and Taxonomy of Clostridia, a Phylogenomic Update.Genome Biol Evol. 2019 Jul 1;11(7):2035-2044. doi: 10.1093/gbe/evz096. Genome Biol Evol. 2019. PMID: 31076745 Free PMC article.
-
Botulinum neurotoxins exploit host digestive proteases to boost their oral toxicity via activating OrfXs/P47.Nat Struct Mol Biol. 2025 May;32(5):864-875. doi: 10.1038/s41594-024-01479-0. Epub 2025 Jan 21. Nat Struct Mol Biol. 2025. PMID: 39838108
-
Clostridial Neurotoxins: Structure, Function and Implications to Other Bacterial Toxins.Microorganisms. 2021 Oct 23;9(11):2206. doi: 10.3390/microorganisms9112206. Microorganisms. 2021. PMID: 34835332 Free PMC article. Review.
-
The Distinctive Evolution of orfX Clostridium parabotulinum Strains and Their Botulinum Neurotoxin Type A and F Gene Clusters Is Influenced by Environmental Factors and Gene Interactions via Mobile Genetic Elements.Front Microbiol. 2021 Feb 26;12:566908. doi: 10.3389/fmicb.2021.566908. eCollection 2021. Front Microbiol. 2021. PMID: 33716993 Free PMC article.
References
-
- Adams PD, Afonine PV, Bunkóczi G, Chen VB, Davis IW, Echols N, Headd JJ, Hung L-W, Kapral GJ, Grosse-Kunstleve RW, McCoy AJ, Moriarty NW, Oeffner R, Read RJ, Richardson DC, Richardson JS, Terwilliger TC, Zwart PH. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D. Biol. Crystallogr. 2010;66:213–21. - PMC - PubMed
-
- Alva V, Lupas AN. The TULIP superfamily of eukaryotic lipid-binding proteins as a mediator of lipid sensing and transport. Biochim. Biophys. Acta - Mol. Cell Biol. Lipids. 2016;1861:913–923. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

