Tracking evolution of aromatase inhibitor resistance with circulating tumour DNA analysis in metastatic breast cancer
- PMID: 29045530
- PMCID: PMC6264798
- DOI: 10.1093/annonc/mdx483
Tracking evolution of aromatase inhibitor resistance with circulating tumour DNA analysis in metastatic breast cancer
Abstract
Background: Selection of resistance mutations may play a major role in the development of endocrine resistance. ESR1 mutations are rare in primary breast cancer but have high prevalence in patients treated with aromatase inhibitors (AI) for advanced breast cancer. We investigated the evolution of genetic resistance to the first-line AI therapy using sequential ctDNA sampling in patients with advanced breast cancer.
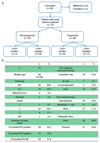
Patients and methods: Eighty-three patients on the first-line AI therapy for metastatic breast cancer were enrolled in a prospective study. Plasma samples were collected every 3 months to disease progression and ctDNA analysed by digital droplet PCR and enhanced tagged-amplicon sequencing (eTAm-Seq). Mutations identified in progression samples by sequencing were tracked back through samples before progression to study the evolution of mutations on therapy. The frequency of novel mutations was validated in an independent cohort of available baseline plasma samples in the Study of Faslodex versus Exemestane with or without Arimidex (SoFEA) trial, which enrolled patients with prior sensitivity to AI.
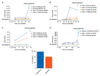
Results: Of the 39 patients who progressed on the first-line AI, 56.4% (22/39) had ESR1 mutations detectable at progression, which were polyclonal in 40.9% (9/22) patients. In serial tracking, ESR1 mutations were detectable median 6.7 months (95% confidence interval 3.7-NA) before clinical progression. Utilising eTAm-Seq ctDNA sequencing of progression plasma, ESR1 mutations were demonstrated to be sub-clonal in 72.2% (13/18) patients. Mutations in RAS genes were identified in 15.4% (6/39) of progressing patients (4 KRAS, 1 HRAS, 1 NRAS). In SoFEA, KRAS mutations were detected in 21.2% (24/113) patients although there was no evidence that KRAS mutation status was prognostic for progression free or overall survival.
Conclusions: Cancers progressing on the first-line AI show high levels of genetic heterogeneity, with frequent sub-clonal mutations. Sub-clonal KRAS mutations are found at high frequency. The genetic diversity of AI resistant cancers may limit subsequent targeted therapy approaches.
Keywords: ESR1; KRAS; breast cancer; ctDNA.
© The Author 2017. Published by Oxford University Press on behalf of the European Society for Medical Oncology. All rights reserved. For permissions, please email: journals.permissions@oup.com.
Figures




References
-
- Misale S, Di Nicolantonio F, Sartore-Bianchi A, Siena S, Bardelli A. Resistance to anti-EGFR therapy in colorectal cancer: from heterogeneity to convergent evolution. Cancer Discov. 2014;4(11):1269–80. - PubMed
-
- Remon J, Caramella C, Jovelet C, Lacroix L, Lawson A, Smalley S, et al. Osimertinib benefit in EGFR-mutant NSCLC patients with T790M-mutation detected by circulating tumour DNA. Ann Oncol. 2017;28(4):784–90. - PubMed
-
- Kobayashi S, Boggon TJ, Dayaram T, Janne PA, Kocher O, Meyerson M, et al. EGFR mutation and resistance of non-small-cell lung cancer to gefitinib. N Engl J Med. 2005;352(8):786–92. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials
Miscellaneous

