MicroRNA gga-miR-130b Suppresses Infectious Bursal Disease Virus Replication via Targeting of the Viral Genome and Cellular Suppressors of Cytokine Signaling 5
- PMID: 29046449
- PMCID: PMC5730774
- DOI: 10.1128/JVI.01646-17
MicroRNA gga-miR-130b Suppresses Infectious Bursal Disease Virus Replication via Targeting of the Viral Genome and Cellular Suppressors of Cytokine Signaling 5
Abstract
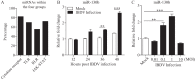
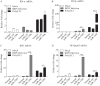
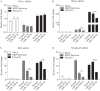
MicroRNAs (miRNAs) are small noncoding RNAs that regulate gene expression posttranscriptionally through silencing or degrading their targets, thus playing important roles in the immune response. However, the role of miRNAs in the host response against infectious bursal disease virus (IBDV) infection is not clear. In this study, we show that the expression of a series of miRNAs was significantly altered in DF-1 cells after IBDV infection. We found that the miRNA gga-miR-130b inhibited IBDV replication via targeting the specific sequence of IBDV segment A and enhanced the expression of beta interferon (IFN-β) by targeting suppressors of cytokine signaling 5 (SOCS5) in host cells. These findings indicate that gga-miR-130b-3p plays a crucial role in host defense against IBDV infection.IMPORTANCE This work shows that gga-miR-130b suppresses IBDV replication via directly targeting the viral genome and cellular SOCS5, the negative regulator for type I interferon expression, revealing the mechanism underlying gga-miR-130-induced inhibition of IBDV replication. This information will be helpful for the understanding of how host cells combat pathogenic infection by self-encoded small RNA and furthers our knowledge of the role of microRNAs in the cell response to viral infection.
Keywords: IBDV; SOCS; microRNA; type I IFN.
Copyright © 2017 American Society for Microbiology.
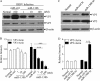
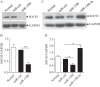
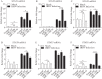
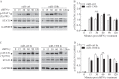
Figures
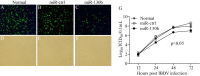
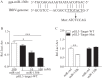
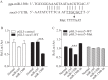
Similar articles
-
Epigenetic Upregulation of Chicken MicroRNA-16-5p Expression in DF-1 Cells following Infection with Infectious Bursal Disease Virus (IBDV) Enhances IBDV-Induced Apoptosis and Viral Replication.J Virol. 2020 Jan 6;94(2):e01724-19. doi: 10.1128/JVI.01724-19. Print 2020 Jan 6. J Virol. 2020. PMID: 31694944 Free PMC article.
-
Role of MicroRNAs in Host Defense against Infectious Bursal Disease Virus (IBDV) Infection: A Hidden Front Line.Viruses. 2020 May 14;12(5):543. doi: 10.3390/v12050543. Viruses. 2020. PMID: 32423052 Free PMC article. Review.
-
gga-miR-454 suppresses infectious bursal disease virus (IBDV) replication via directly targeting IBDV genomic segment B and cellular Suppressors of Cytokine Signaling 6 (SOCS6).Virus Res. 2018 Jul 2;252:29-40. doi: 10.1016/j.virusres.2018.05.015. Epub 2018 May 16. Virus Res. 2018. PMID: 29777734
-
gga-miR-155 Enhances Type I Interferon Expression and Suppresses Infectious Burse Disease Virus Replication via Targeting SOCS1 and TANK.Front Cell Infect Microbiol. 2018 Mar 7;8:55. doi: 10.3389/fcimb.2018.00055. eCollection 2018. Front Cell Infect Microbiol. 2018. PMID: 29564226 Free PMC article.
-
Regulation of the MIR155 host gene in physiological and pathological processes.Gene. 2013 Dec 10;532(1):1-12. doi: 10.1016/j.gene.2012.12.009. Epub 2012 Dec 14. Gene. 2013. PMID: 23246696 Review.
Cited by
-
Integrated mRNA and miRNA profiling in NIH/3T3 cells in response to bovine papillomavirus E6 gene expression.PeerJ. 2019 Aug 2;7:e7442. doi: 10.7717/peerj.7442. eCollection 2019. PeerJ. 2019. PMID: 31396463 Free PMC article.
-
The Influence of Host miRNA Binding to RNA Within RNA Viruses on Virus Multiplication.Front Cell Infect Microbiol. 2022 Apr 21;12:802149. doi: 10.3389/fcimb.2022.802149. eCollection 2022. Front Cell Infect Microbiol. 2022. PMID: 35531344 Free PMC article. Review.
-
The Roles of MicroRNAs (miRNAs) in Avian Response to Viral Infection and Pathogenesis of Avian Immunosuppressive Diseases.Int J Mol Sci. 2019 Nov 1;20(21):5454. doi: 10.3390/ijms20215454. Int J Mol Sci. 2019. PMID: 31683847 Free PMC article. Review.
-
Epigenetic Upregulation of Chicken MicroRNA-16-5p Expression in DF-1 Cells following Infection with Infectious Bursal Disease Virus (IBDV) Enhances IBDV-Induced Apoptosis and Viral Replication.J Virol. 2020 Jan 6;94(2):e01724-19. doi: 10.1128/JVI.01724-19. Print 2020 Jan 6. J Virol. 2020. PMID: 31694944 Free PMC article.
-
Role of MicroRNAs in Host Defense against Infectious Bursal Disease Virus (IBDV) Infection: A Hidden Front Line.Viruses. 2020 May 14;12(5):543. doi: 10.3390/v12050543. Viruses. 2020. PMID: 32423052 Free PMC article. Review.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

