Distinct Roles of Cellular ESCRT-I and ESCRT-III Proteins in Efficient Entry and Egress of Budded Virions of Autographa californica Multiple Nucleopolyhedrovirus
- PMID: 29046462
- PMCID: PMC5730794
- DOI: 10.1128/JVI.01636-17
Distinct Roles of Cellular ESCRT-I and ESCRT-III Proteins in Efficient Entry and Egress of Budded Virions of Autographa californica Multiple Nucleopolyhedrovirus
Abstract
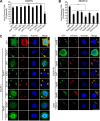
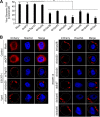
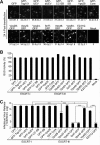
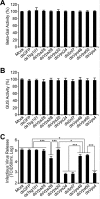
The endosomal sorting complex required for transport (ESCRT) machinery is necessary for budding of many enveloped viruses. Recently, it was demonstrated that Vps4, the key regulator for recycling of the ESCRT-III complex, is required for efficient infection by the baculovirus Autographa californica multiple nucleopolyhedrovirus (AcMNPV). However, ESCRT assembly, regulation, and function are complex, and little is known regarding the details of participation of specific ESCRT complexes in AcMNPV infection. In this study, the core components of ESCRT-I (Tsg101 and Vps28) and ESCRT-III (Vps2B, Vps20, Vps24, Snf7, Vps46, and Vps60) were cloned from Spodoptera frugiperda Using a viral complementation system and RNA interference (RNAi) assays, we found that ESCRT-I and ESCRT-III complexes are required for efficient entry of AcMNPV into insect cells. In cells knocking down or overexpressing dominant negative (DN) forms of the components of ESCRT-I and ESCRT-III complexes, entering virions were partially trapped within the cytosol. To examine only egress, cells were transfected with the double-stranded RNA (dsRNA) targeting an individual ESCRT-I or ESCRT-III gene and viral bacmid DNA or viral bacmid DNA that expressed DN forms of ESCRT-I and ESCRT-III components. We found that ESCRT-III components (but not ESCRT-I components) are required for efficient nuclear egress of progeny nucleocapsids. In addition, we found that several baculovirus core or conserved proteins (Ac11, Ac76, Ac78, GP41, Ac93, Ac103, Ac142, and Ac146) interact with Vps4 and components of ESCRT-III. We propose that these viral proteins may form an "egress complex" that is involved in recruiting ESCRT-III components to a virus egress domain on the nuclear membrane.IMPORTANCE The ESCRT system is hijacked by many enveloped viruses to mediate budding and release. Recently, it was found that Vps4, the key regulator of the cellular ESCRT machinery, is necessary for efficient entry and egress of Autographa californica multiple nucleopolyhedrovirus (AcMNPV). However, little is known about the roles of specific ESCRT complexes in AcMNPV infection. In this study, we demonstrated that ESCRT-I and ESCRT-III complexes are required for efficient entry of AcMNPV into insect cells. The components of ESCRT-III (but not ESCRT-I) are also necessary for efficient nuclear egress of progeny nucleocapsids. Several baculovirus core or conserved proteins were found to interact with Vps4 and components of ESCRT-III, and these interactions may suggest the formation of an "egress complex" involved in the nuclear release or transport of viral nucleocapsids.
Keywords: AcMNPV; ESCRT-I; ESCRT-III; baculovirus; virus entry and egress.
Copyright © 2017 American Society for Microbiology.
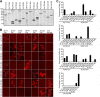
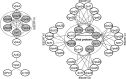
Figures

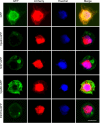
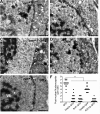
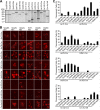
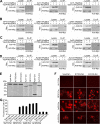

Similar articles
-
Multifaceted interactions between host ESCRT-III and budded virus-related proteins involved in entry and egress of the baculovirus Autographa californica multiple nucleopolyhedrovirus.J Virol. 2024 Feb 20;98(2):e0190023. doi: 10.1128/jvi.01900-23. Epub 2024 Jan 30. J Virol. 2024. PMID: 38289107 Free PMC article.
-
Roles of Cellular NSF Protein in Entry and Nuclear Egress of Budded Virions of Autographa californica Multiple Nucleopolyhedrovirus.J Virol. 2017 Sep 27;91(20):e01111-17. doi: 10.1128/JVI.01111-17. Print 2017 Oct 15. J Virol. 2017. PMID: 28747507 Free PMC article.
-
Cellular VPS4 is required for efficient entry and egress of budded virions of Autographa californica multiple nucleopolyhedrovirus.J Virol. 2012 Jan;86(1):459-72. doi: 10.1128/JVI.06049-11. Epub 2011 Nov 9. J Virol. 2012. PMID: 22072775 Free PMC article.
-
[Cellular ESCRT complex and its roles in enveloped viruses budding].Sheng Wu Gong Cheng Xue Bao. 2012 Sep;28(9):1031-7. Sheng Wu Gong Cheng Xue Bao. 2012. PMID: 23289305 Review. Chinese.
-
Baculovirus Entry and Egress from Insect Cells.Annu Rev Virol. 2018 Sep 29;5(1):113-139. doi: 10.1146/annurev-virology-092917-043356. Epub 2018 Jul 13. Annu Rev Virol. 2018. PMID: 30004832 Review.
Cited by
-
The Functional Oligomeric State of Tegument Protein GP41 Is Essential for Baculovirus Budded Virion and Occlusion-Derived Virion Assembly.J Virol. 2018 May 29;92(12):e02083-17. doi: 10.1128/JVI.02083-17. Print 2018 Jun 15. J Virol. 2018. PMID: 29643237 Free PMC article.
-
Identification and characterization of coiled-coil motifs across Autographa californica multiple nucleopolyhedrovirus genome.Heliyon. 2022 Sep 10;8(9):e10588. doi: 10.1016/j.heliyon.2022.e10588. eCollection 2022 Sep. Heliyon. 2022. PMID: 36132175 Free PMC article.
-
Host AAA+ ATPase TER94 Plays Critical Roles in Building the Baculovirus Viral Replication Factory and Virion Morphogenesis.J Virol. 2020 Feb 28;94(6):e01674-19. doi: 10.1128/JVI.01674-19. Print 2020 Feb 28. J Virol. 2020. PMID: 31896597 Free PMC article.
-
Improvement of protein production in baculovirus expression vector system by removing a total of 10 kb of nonessential fragments from Autographa californica multiple nucleopolyhedrovirus genome.Front Microbiol. 2023 Apr 13;14:1171500. doi: 10.3389/fmicb.2023.1171500. eCollection 2023. Front Microbiol. 2023. PMID: 37125202 Free PMC article.
-
Autographa californica multiple nucleopolyhedrovirus ac106 is required for the nuclear egress of nucleocapsids and intranuclear microvesicle formation.J Virol. 2024 Nov 19;98(11):e0113524. doi: 10.1128/jvi.01135-24. Epub 2024 Oct 21. J Virol. 2024. PMID: 39431847 Free PMC article.
References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

