An integrative in-silico approach for therapeutic target identification in the human pathogen Corynebacterium diphtheriae
- PMID: 29049350
- PMCID: PMC5648181
- DOI: 10.1371/journal.pone.0186401
An integrative in-silico approach for therapeutic target identification in the human pathogen Corynebacterium diphtheriae
Abstract
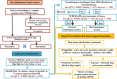
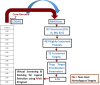
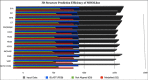
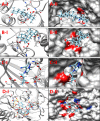
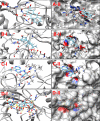
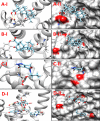
Corynebacterium diphtheriae (Cd) is a Gram-positive human pathogen responsible for diphtheria infection and once regarded for high mortalities worldwide. The fatality gradually decreased with improved living standards and further alleviated when many immunization programs were introduced. However, numerous drug-resistant strains emerged recently that consequently decreased the efficacy of current therapeutics and vaccines, thereby obliging the scientific community to start investigating new therapeutic targets in pathogenic microorganisms. In this study, our contributions include the prediction of modelome of 13 C. diphtheriae strains, using the MHOLline workflow. A set of 463 conserved proteins were identified by combining the results of pangenomics based core-genome and core-modelome analyses. Further, using subtractive proteomics and modelomics approaches for target identification, a set of 23 proteins was selected as essential for the bacteria. Considering human as a host, eight of these proteins (glpX, nusB, rpsH, hisE, smpB, bioB, DIP1084, and DIP0983) were considered as essential and non-host homologs, and have been subjected to virtual screening using four different compound libraries (extracted from the ZINC database, plant-derived natural compounds and Di-terpenoid Iso-steviol derivatives). The proposed ligand molecules showed favorable interactions, lowered energy values and high complementarity with the predicted targets. Our proposed approach expedites the selection of C. diphtheriae putative proteins for broad-spectrum development of novel drugs and vaccines, owing to the fact that some of these targets have already been identified and validated in other organisms.
Conflict of interest statement
Figures






Similar articles
-
Proteome scale comparative modeling for conserved drug and vaccine targets identification in Corynebacterium pseudotuberculosis.BMC Genomics. 2014;15 Suppl 7(Suppl 7):S3. doi: 10.1186/1471-2164-15-S7-S3. Epub 2014 Oct 27. BMC Genomics. 2014. PMID: 25573232 Free PMC article.
-
The Druggable Pocketome of Corynebacterium diphtheriae: A New Approach for in silico Putative Druggable Targets.Front Genet. 2018 Feb 13;9:44. doi: 10.3389/fgene.2018.00044. eCollection 2018. Front Genet. 2018. PMID: 29487617 Free PMC article.
-
Conserved host-pathogen PPIs. Globally conserved inter-species bacterial PPIs based conserved host-pathogen interactome derived novel target in C. pseudotuberculosis, C. diphtheriae, M. tuberculosis, C. ulcerans, Y. pestis, and E. coli targeted by Piper betel compounds.Integr Biol (Camb). 2013 Mar;5(3):495-509. doi: 10.1039/c2ib20206a. Integr Biol (Camb). 2013. PMID: 23288366
-
Evolution, epidemiology and diversity of Corynebacterium diphtheriae: New perspectives on an old foe.Infect Genet Evol. 2016 Sep;43:364-70. doi: 10.1016/j.meegid.2016.06.024. Epub 2016 Jun 9. Infect Genet Evol. 2016. PMID: 27291708 Review.
-
Corynebacterium diphtheriae: genome diversity, population structure and genotyping perspectives.Infect Genet Evol. 2009 Jan;9(1):1-15. doi: 10.1016/j.meegid.2008.09.011. Epub 2008 Oct 19. Infect Genet Evol. 2009. PMID: 19007916 Review.
Cited by
-
Neuroprotective Potential of Synthetic Mono-Carbonyl Curcumin Analogs Assessed by Molecular Docking Studies.Molecules. 2021 Nov 26;26(23):7168. doi: 10.3390/molecules26237168. Molecules. 2021. PMID: 34885751 Free PMC article.
-
Designing a multi-epitope vaccine against Shigella dysenteriae using immuno-informatics approach.Front Genet. 2024 May 17;15:1361610. doi: 10.3389/fgene.2024.1361610. eCollection 2024. Front Genet. 2024. PMID: 38826807 Free PMC article.
-
An Integrated Database of Small RNAs and Their Interplay With Transcriptional Gene Regulatory Networks in Corynebacteria.Front Microbiol. 2021 Jun 17;12:656435. doi: 10.3389/fmicb.2021.656435. eCollection 2021. Front Microbiol. 2021. PMID: 34220744 Free PMC article.
-
Proteomics of Toxigenic Corynebacteria.Proteomes. 2023 Dec 30;12(1):2. doi: 10.3390/proteomes12010002. Proteomes. 2023. PMID: 38250813 Free PMC article. Review.
-
Genomic landscape of the emerging XDR Salmonella Typhi for mining druggable targets clpP, hisH, folP and gpmI and screening of novel TCM inhibitors, molecular docking and simulation analyses.BMC Microbiol. 2023 Jan 21;23(1):25. doi: 10.1186/s12866-023-02756-6. BMC Microbiol. 2023. PMID: 36681806 Free PMC article.
References
-
- Goodfellow M, Kämpfer P,. Busse HJ, Trujillo M, Suzuki KI, Ludwig W. Whitman Bergey’s manual of systematic bacteriology: Springer; 2012.
-
- Hodes HL. Diphtheria. Pediatr Clin North Am. 1979;26(2):445–59. . - PubMed
-
- Wagner KS, White JM, Crowcroft NS, De Martin S, Mann G, Efstratiou A. Diphtheria in the United Kingdom, 1986–2008: the increasing role of Corynebacterium ulcerans. Epidemiol Infect. 2010;138(11):1519–30. doi: 10.1017/S0950268810001895 . - DOI - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

