High-frequency, low-coverage "false positives" mutations may be true in GS Junior sequencing studies
- PMID: 29062110
- PMCID: PMC5653793
- DOI: 10.1038/s41598-017-13116-6
High-frequency, low-coverage "false positives" mutations may be true in GS Junior sequencing studies
Abstract
The GS Junior sequencer provides simplified procedures for library preparation and data processing. Errors in pyrosequencing generate some biases during library construction and emulsion PCR amplification. False-positive mutations are identified by related characteristics described in the manufacturer's manual, and some detected mutations may have 'borderline' characteristics when they are detected in few reads or at low frequency. Among these mutations, however, some may be true positives. This study aimed to improve the accuracy of identifying true positives among mutations with borderline false-positive characteristics detected with GS Junior sequencing. Mutations with the borderline features were tested for validity with Sanger sequencing. We examined 10 mutations detected in coverages <20-fold at frequencies >30% (group A) and 16 mutations detected in coverages >20-fold at frequencies < 30% (group B). In group A, two mutations were not confirmed, and two mutations with 100% frequency were confirmed as heterozygous alleles. No mutation in group B was confirmed. The two groups had significantly different false-positive prevalences (p = 0.001). These results suggest that mutations detected at frequencies less than 30% can be confidently identified as false-positives but that mutations detected at frequencies over 30%, despite coverages less than 20-fold, should be verified with Sanger sequencing.
Conflict of interest statement
The authors declare that they have no competing interests.
Figures
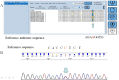
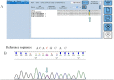
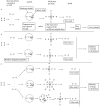
References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

