Single-cell heterogeneity and cell-cycle-related viral gene bursts in the human leukaemia virus HTLV-1
- PMID: 29062917
- PMCID: PMC5645716
- DOI: 10.12688/wellcomeopenres.12469.2
Single-cell heterogeneity and cell-cycle-related viral gene bursts in the human leukaemia virus HTLV-1
Abstract
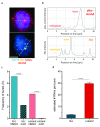
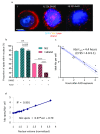
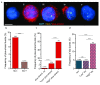
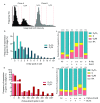
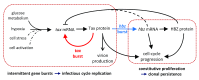
Background: The human leukaemia virus HTLV-1 expresses essential accessory genes that manipulate the expression, splicing and transport of viral mRNAs. Two of these genes, tax and hbz, also promote proliferation of the infected cell, and both genes are thought to contribute to oncogenesis in adult T-cell leukaemia/lymphoma. The regulation of HTLV-1 proviral latency is not understood. tax, on the proviral plus strand, is usually silent in freshly-isolated cells, whereas the minus-strand-encoded hbz gene is persistently expressed at a low level. However, the persistently activated host immune response to Tax indicates frequent expression of tax in vivo. Methods: We used single-molecule RNA-FISH to quantify the expression of HTLV-1 transcripts at the single-cell level in a total of >19,000 cells from five T-cell clones, naturally infected with HTLV-1, isolated by limiting dilution from peripheral blood of HTLV-1-infected subjects. Results: We found strong heterogeneity both within and between clones in the expression of the proviral plus-strand (detected by hybridization to the tax gene) and the minus-strand ( hbz gene). Both genes are transcribed in bursts; tax expression is enhanced in the absence of hbz, while hbz expression increased in cells with high tax expression. Surprisingly, we found that hbz expression is strongly associated with the S and G 2/M phases of the cell cycle, independent of tax expression. Contrary to current belief, hbz is not expressed in all cells at all times, even within one clone. In hbz-positive cells, the abundance of hbz transcripts showed a very strong positive linear correlation with nuclear volume. Conclusions: The occurrence of intense, intermittent plus-strand gene bursts in independent primary HTLV-1-infected T-cell clones from unrelated individuals strongly suggests that the HTLV-1 plus-strand is expressed in bursts in vivo. Our results offer an explanation for the paradoxical correlations observed between the host immune response and HTLV-1 transcription.
Keywords: HTLV-; RNA-FISH; gene burst; latency; retrovirus; smFISH; transcription.
Conflict of interest statement
Competing interests: No competing interests were disclosed.
Figures

References
-
- Hanon E, Hall S, Taylor GP, et al. : Abundant tax protein expression in CD4+ T cells infected with human T-cell lymphotropic virus type I (HTLV-I) is prevented by cytotoxic T lymphocytes. Blood. 2000;95(4):1386–1392. - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

