Deep transcriptome annotation enables the discovery and functional characterization of cryptic small proteins
- PMID: 29083303
- PMCID: PMC5703645
- DOI: 10.7554/eLife.27860
Deep transcriptome annotation enables the discovery and functional characterization of cryptic small proteins
Abstract
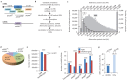
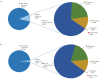
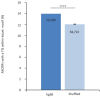
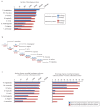
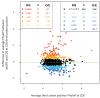
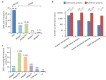
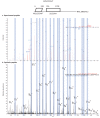
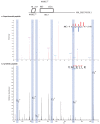
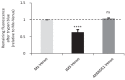
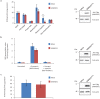
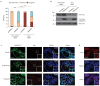
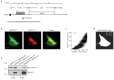
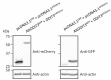
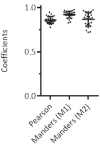
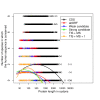
Recent functional, proteomic and ribosome profiling studies in eukaryotes have concurrently demonstrated the translation of alternative open-reading frames (altORFs) in addition to annotated protein coding sequences (CDSs). We show that a large number of small proteins could in fact be coded by these altORFs. The putative alternative proteins translated from altORFs have orthologs in many species and contain functional domains. Evolutionary analyses indicate that altORFs often show more extreme conservation patterns than their CDSs. Thousands of alternative proteins are detected in proteomic datasets by reanalysis using a database containing predicted alternative proteins. This is illustrated with specific examples, including altMiD51, a 70 amino acid mitochondrial fission-promoting protein encoded in MiD51/Mief1/SMCR7L, a gene encoding an annotated protein promoting mitochondrial fission. Our results suggest that many genes are multicoding genes and code for a large protein and one or several small proteins.
Keywords: alternative translation; biochemistry; computational biology; human; open reading frames; small proteins; systems biology; translation; translation initiation sites.
Conflict of interest statement
No competing interests declared.
Figures



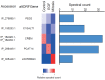
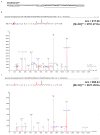
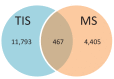
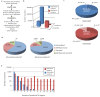
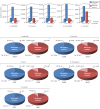
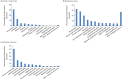
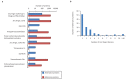
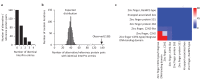

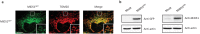


Similar articles
-
The Protein Coded by a Short Open Reading Frame, Not by the Annotated Coding Sequence, Is the Main Gene Product of the Dual-Coding Gene MIEF1.Mol Cell Proteomics. 2018 Dec;17(12):2402-2411. doi: 10.1074/mcp.RA118.000593. Epub 2018 Sep 4. Mol Cell Proteomics. 2018. PMID: 30181344 Free PMC article.
-
Direct detection of alternative open reading frames translation products in human significantly expands the proteome.PLoS One. 2013 Aug 12;8(8):e70698. doi: 10.1371/journal.pone.0070698. eCollection 2013. PLoS One. 2013. PMID: 23950983 Free PMC article.
-
Evaluation of Eukaryotic mRNA Coding Potential.Methods Mol Biol. 2025;2859:319-331. doi: 10.1007/978-1-0716-4152-1_18. Methods Mol Biol. 2025. PMID: 39436610
-
Small Proteins Encoded by Unannotated ORFs are Rising Stars of the Proteome, Confirming Shortcomings in Genome Annotations and Current Vision of an mRNA.Proteomics. 2018 May;18(10):e1700058. doi: 10.1002/pmic.201700058. Epub 2017 Oct 11. Proteomics. 2018. PMID: 28627015 Review.
-
Translation of Small Open Reading Frames: Roles in Regulation and Evolutionary Innovation.Trends Genet. 2019 Mar;35(3):186-198. doi: 10.1016/j.tig.2018.12.003. Epub 2018 Dec 31. Trends Genet. 2019. PMID: 30606460 Review.
Cited by
-
Comparative Proteomic Profiling of Unannotated Microproteins and Alternative Proteins in Human Cell Lines.J Proteome Res. 2020 Aug 7;19(8):3418-3426. doi: 10.1021/acs.jproteome.0c00254. Epub 2020 Jun 3. J Proteome Res. 2020. PMID: 32449352 Free PMC article.
-
Principles, mechanisms, and biological implications of translation termination-reinitiation.RNA. 2023 Jul;29(7):865-884. doi: 10.1261/rna.079375.122. Epub 2023 Apr 6. RNA. 2023. PMID: 37024263 Free PMC article.
-
The FUS gene is dual-coding with both proteins contributing to FUS-mediated toxicity.EMBO Rep. 2021 Jan 7;22(1):e50640. doi: 10.15252/embr.202050640. Epub 2020 Nov 23. EMBO Rep. 2021. PMID: 33226175 Free PMC article.
-
Microproteins in Metabolism.Cells. 2025 Jun 7;14(12):859. doi: 10.3390/cells14120859. Cells. 2025. PMID: 40558487 Free PMC article. Review.
-
Protein evidence of unannotated ORFs in Drosophila reveals diversity in the evolution and properties of young proteins.Elife. 2022 Sep 30;11:e78772. doi: 10.7554/eLife.78772. Elife. 2022. PMID: 36178469 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

