A comprehensive approach to identification of pathogenic FANCA variants in Fanconi anemia patients and their families
- PMID: 29098742
- PMCID: PMC5762269
- DOI: 10.1002/humu.23366
A comprehensive approach to identification of pathogenic FANCA variants in Fanconi anemia patients and their families
Abstract
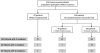
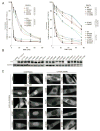
Fanconi anemia (FA) is a rare recessive DNA repair deficiency resulting from mutations in one of at least 22 genes. Two-thirds of FA families harbor mutations in FANCA. To genotype patients in the International Fanconi Anemia Registry (IFAR) we employed multiple methodologies, screening 216 families for FANCA mutations. We describe identification of 57 large deletions and 261 sequence variants, in 159 families. All but seven families harbored distinct combinations of two mutations demonstrating high heterogeneity. Pathogenicity of the 18 novel missense variants was analyzed functionally by determining the ability of the mutant cDNA to improve the survival of a FANCA-null cell line when treated with MMC. Overexpressed pathogenic missense variants were found to reside in the cytoplasm, and nonpathogenic in the nucleus. RNA analysis demonstrated that two variants (c.522G > C and c.1565A > G), predicted to encode missense variants, which were determined to be nonpathogenic by a functional assay, caused skipping of exons 5 and 16, respectively, and are most likely pathogenic. We report 48 novel FANCA sequence variants. Defining both variants in a large patient cohort is a major step toward cataloging all FANCA variants, and permitting studies of genotype-phenotype correlations.
Keywords: FANCA; Fanconi anemia; functional assay; pathogenic mutations; recessive disorder.
© Published 2017. This article is a U.S. Government work and is in the public domain in the USA.
Conflict of interest statement
Figures



References
-
- Adachi D, Oda T, Yagasaki H, Nakasato K, Taniguchi T, D'Andrea AD, Asano S, Yamashita T. Heterogeneous activation of the Fanconi anemia pathway by patient-derived FANCA mutants. Hum Mol Genet. 2002;11(25):3125–34. - PubMed
-
- Alter BP, Giri N. Thinking of VACTERL-H? Rule out Fanconi Anemia according to PHENOS. Am J Med Genet A. 2016;170(6):1520–4. - PubMed
-
- Ameziane N, Errami A, Leveille F, Fontaine C, de Vries Y, van Spaendonk RM, de Winter JP, Pals G, Joenje H. Genetic subtyping of Fanconi anemia by comprehensive mutation screening. Hum Mutat. 2008;29(1):159–66. - PubMed
-
- Amouri A, Talmoudi F, Messaoud O, d'Enghien CD, Rekaya MB, Allegui I, Azaiez H, Kefi R, Abdelhak A, Meseddi SH, Torjemane L, Ouederni M, Mellouli F, Abid HB, Aissaoui L, Bejaoui M, Othmen TB, Lyonnet DS, Soulier J, Hachicha M, Dellagi K, Abdelhak S, Fanconi T. High frequency of exon 15 deletion in the FANCA gene in Tunisian patients affected with Fanconi anemia disease: implication for diagnosis. Mol Genet Genomic Med. 2014;2(2):160–5. - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

