Mutant p53 partners in crime
- PMID: 29099488
- PMCID: PMC5729539
- DOI: 10.1038/cdd.2017.185
Mutant p53 partners in crime
Abstract
Mutant p53 proteins impart changes in cellular behavior and function through interactions with proteins that alter gene expression. The milieu of intracellular proteins available to interact with mutant p53 is context specific and changes with disease, cell type, and environmental conditions. Varying conformations of mutant p53 largely dictate protein-protein interactions as different point mutations within protein-coding regions greatly alter the extent and array of gain-of-function (GOF) activities. Given such variables, how can knowledge regarding p53 missense mutations be translated into predicting or altering biologic activity for therapy? How may knowledge regarding mutant p53 functions within certain disease contexts be harnessed to blunt or ablate mutant p53 GOF for therapy? In this article, we review known proteins that interact with mutant p53 and result in the activation of genes that contribute to p53 GOF with particular emphasis on context dependency and an evolving appreciation of GOF mechanisms.
Conflict of interest statement
The authors declare no conflict of interest.
Figures
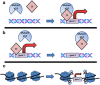
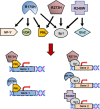
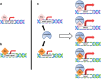
References
-
- Willis A, Jung EJ, Wakefield T, Chen X. Mutant p53 exerts a dominant negative effect by preventing wild-type p53 from binding to the promoter of its target genes. Oncogene 2004; 23: 2330–2338. - PubMed
-
- Lang GA, Iwakuma T, Suh YA, Liu G, Rao VA, Parant JM et al. Gain of function of a p53 hot spot mutation in a mouse model of Li-Fraumeni syndrome. Cell 2004; 119: 861–872. - PubMed
-
- Milner J, Medcalf EA. Cotranslation of activated mutant p53 with wild type drives the wild-type p53 protein into the mutant conformation. Cell 1991; 65: 765–774. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous

