CRISPR interference-guided multiplex repression of endogenous competing pathway genes for redirecting metabolic flux in Escherichia coli
- PMID: 29100516
- PMCID: PMC5670510
- DOI: 10.1186/s12934-017-0802-x
CRISPR interference-guided multiplex repression of endogenous competing pathway genes for redirecting metabolic flux in Escherichia coli
Abstract
Background: Multiplex control of metabolic pathway genes is essential for maximizing product titers and conversion yields of fuels, chemicals, and pharmaceuticals in metabolic engineering. To achieve this goal, artificial transcriptional regulators, such as clustered regularly interspaced short palindromic repeats (CRISPR) interference (CRISPRi), have been developed to specifically repress genes of interest.
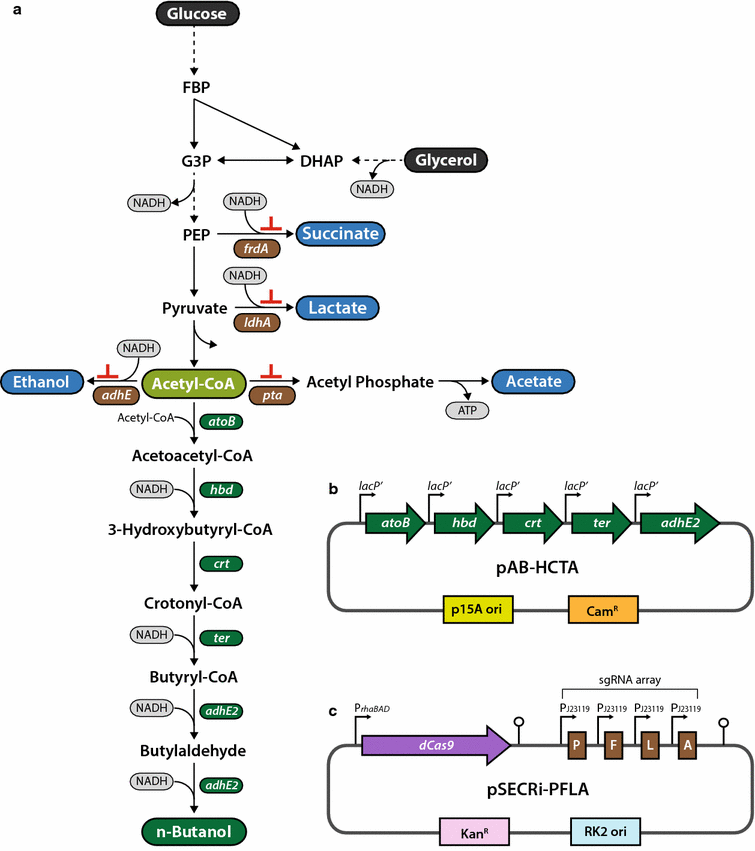
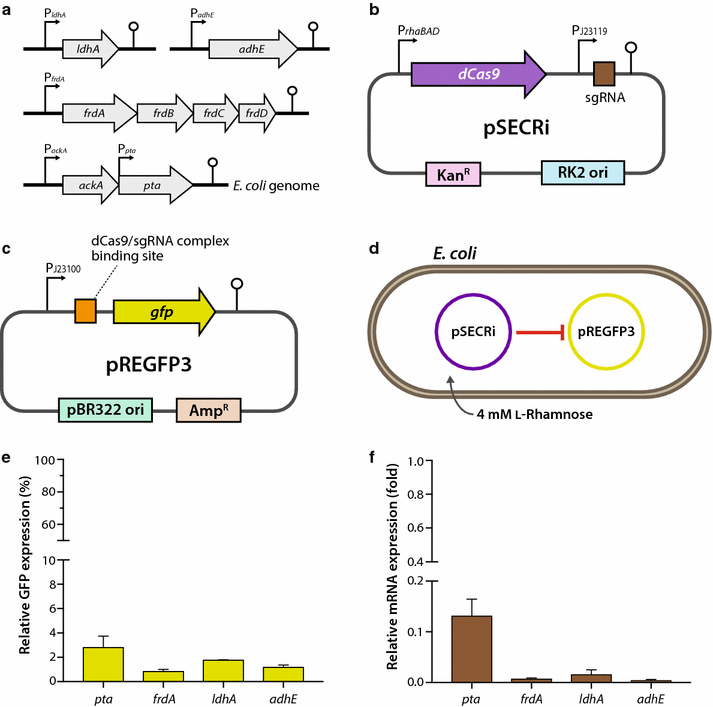
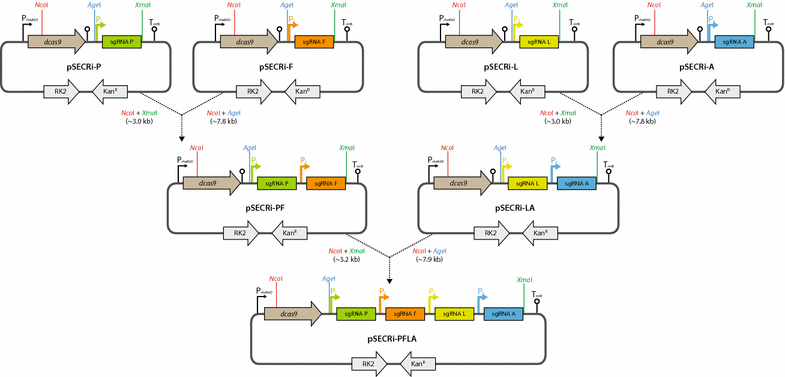
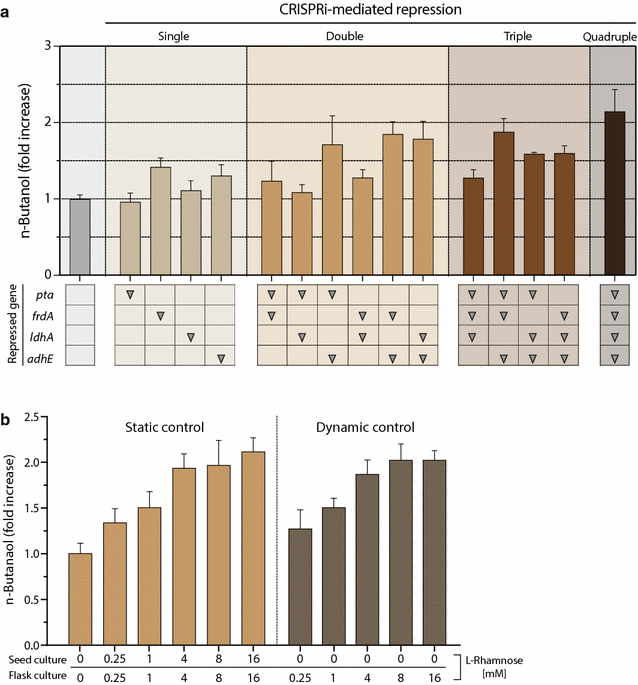
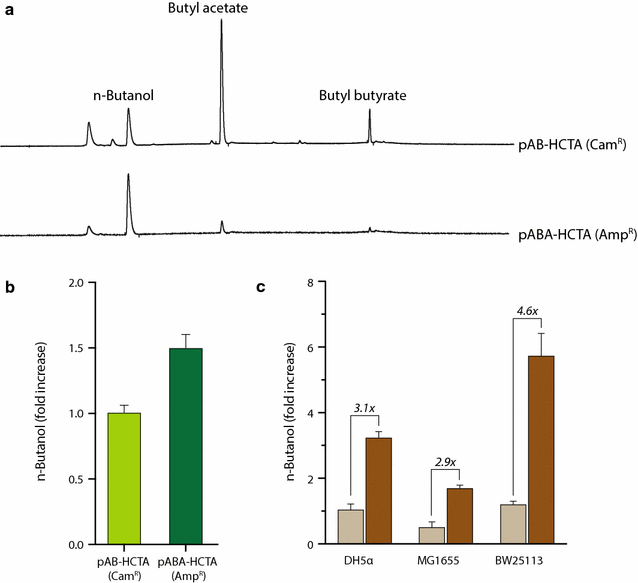
Results: In this study, we deployed a tunable CRISPRi system for multiplex repression of competing pathway genes and, thus, directed carbon flux toward production of molecules of interest in Escherichia coli. The tunable CRISPRi system with an array of sgRNAs successfully repressed four endogenous genes (pta, frdA, ldhA, and adhE) individually and in double, triple, or quadruple combination that are involved in the formation of byproducts (acetate, succinate, lactate, and ethanol) and the consumption of NADH in E. coli. Single-target CRISPRi effectively reduced the amount of each byproduct and, interestingly, pta repression also decreased ethanol production (41%), whereas ldhA repression increased ethanol production (197%). CRISPRi-mediated multiplex repression of competing pathway genes also resulted in simultaneous reductions of acetate, succinate, lactate, and ethanol production in E. coli. Among 15 conditions repressing byproduct-formation genes, we chose the quadruple-target CRISPRi condition to produce n-butanol in E. coli as a case study. When heterologous n-butanol-pathway enzymes were introduced into E. coli simultaneously repressing the expression of the pta, frdA, ldhA, and adhE genes via CRISPRi, n-butanol yield and productivity increased up to 5.4- and 3.2-fold, respectively.
Conclusions: We demonstrated the tunable CRISPRi system to be a robust platform for multiplex modulation of endogenous gene expression that can be used to enhance biosynthetic pathway productivity, with n-butanol as the test case. CRISPRi applications potentially enable the development of microbial "smart cell" factories capable of producing other industrially valuable products.
Keywords: CRISPR interference; Endogenous gene; Escherichia coli; Multiple gene knockdown; n-Butanol.
Figures
Similar articles
-
CRISPR interference-guided balancing of a biosynthetic mevalonate pathway increases terpenoid production.Metab Eng. 2016 Nov;38:228-240. doi: 10.1016/j.ymben.2016.08.006. Epub 2016 Aug 26. Metab Eng. 2016. PMID: 27569599
-
Redirecting Metabolic Flux via Combinatorial Multiplex CRISPRi-Mediated Repression for Isopentenol Production in Escherichia coli.ACS Synth Biol. 2019 Feb 15;8(2):391-402. doi: 10.1021/acssynbio.8b00429. Epub 2019 Feb 5. ACS Synth Biol. 2019. PMID: 30681833
-
dCas12a-mediated CRISPR interference for multiplex gene repression in cyanobacteria for enhanced isobutanol and 3-methyl-1-butanol production.Microb Cell Fact. 2025 May 13;24(1):104. doi: 10.1186/s12934-025-02727-8. Microb Cell Fact. 2025. PMID: 40361099 Free PMC article.
-
CRISPR-Cas9/CRISPRi tools for cell factory construction in E. coli.World J Microbiol Biotechnol. 2020 Jun 25;36(7):96. doi: 10.1007/s11274-020-02872-9. World J Microbiol Biotechnol. 2020. PMID: 32583135 Review.
-
Advances and applications of CRISPR/Cas-mediated interference in Escherichia coli.Eng Microbiol. 2023 Nov 2;4(1):100123. doi: 10.1016/j.engmic.2023.100123. eCollection 2024 Mar. Eng Microbiol. 2023. PMID: 39628789 Free PMC article. Review.
Cited by
-
Repurposing CRISPR-Cas Systems as Genetic Tools for the Enterobacteriales.EcoSal Plus. 2021 Dec 15;9(2):eESP00062020. doi: 10.1128/ecosalplus.ESP-0006-2020. Epub 2021 Jun 14. EcoSal Plus. 2021. PMID: 34125584 Free PMC article. Review.
-
CRISPR-Cas9/Cas12a biotechnology and application in bacteria.Synth Syst Biotechnol. 2018 Oct 3;3(3):135-149. doi: 10.1016/j.synbio.2018.09.004. eCollection 2018 Sep. Synth Syst Biotechnol. 2018. PMID: 30345399 Free PMC article. Review.
-
Regulation of Microbial Metabolic Rates Using CRISPR Interference With Expanded PAM Sequences.Front Microbiol. 2020 Feb 28;11:282. doi: 10.3389/fmicb.2020.00282. eCollection 2020. Front Microbiol. 2020. PMID: 32184769 Free PMC article.
-
Genome-scale target identification in Escherichia coli for high-titer production of free fatty acids.Nat Commun. 2021 Aug 17;12(1):4976. doi: 10.1038/s41467-021-25243-w. Nat Commun. 2021. PMID: 34404790 Free PMC article.
-
Applications of CRISPR/Cas System to Bacterial Metabolic Engineering.Int J Mol Sci. 2018 Apr 5;19(4):1089. doi: 10.3390/ijms19041089. Int J Mol Sci. 2018. PMID: 29621180 Free PMC article. Review.
References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

