The Co-regulation Data Harvester: automating gene annotation starting from a transcriptome database
- PMID: 29104906
- PMCID: PMC5663188
- DOI: 10.1016/j.softx.2017.06.006
The Co-regulation Data Harvester: automating gene annotation starting from a transcriptome database
Abstract
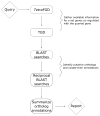
Identifying co-regulated genes provides a useful approach for defining pathway-specific machinery in an organism. To be efficient, this approach relies on thorough genome annotation, a process much slower than genome sequencing per se. Tetrahymena thermophila, a unicellular eukaryote, has been a useful model organism and has a fully sequenced but sparsely annotated genome. One important resource for studying this organism has been an online transcriptomic database. We have developed an automated approach to gene annotation in the context of transcriptome data in T. thermophila, called the Co-regulation Data Harvester (CDH). Beginning with a gene of interest, the CDH identifies co-regulated genes by accessing the Tetrahymena transcriptome database. It then identifies their closely related genes (orthologs) in other organisms by using reciprocal BLAST searches. Finally, it collates the annotations of those orthologs' functions, which provides the user with information to help predict the cellular role of the initial query. The CDH, which is freely available, represents a powerful new tool for analyzing cell biological pathways in Tetrahymena. Moreover, to the extent that genes and pathways are conserved between organisms, the inferences obtained via the CDH should be relevant, and can be explored, in many other systems.
Keywords: Automation; Bioinformatics; Evolution; Protists.
Figures


Similar articles
-
Tetrahymena functional genomics database (TetraFGD): an integrated resource for Tetrahymena functional genomics.Database (Oxford). 2013 Mar 12;2013:bat008. doi: 10.1093/database/bat008. Print 2013. Database (Oxford). 2013. PMID: 23482072 Free PMC article.
-
Tetrahymena Comparative Genomics Database (TCGD): a community resource for Tetrahymena.Database (Oxford). 2019 Jan 1;2019:baz029. doi: 10.1093/database/baz029. Database (Oxford). 2019. PMID: 30810209 Free PMC article.
-
Tetrahymena Genome Database (TGD): a new genomic resource for Tetrahymena thermophila research.Nucleic Acids Res. 2006 Jan 1;34(Database issue):D500-3. doi: 10.1093/nar/gkj054. Nucleic Acids Res. 2006. PMID: 16381920 Free PMC article.
-
Getting the most out of parasitic helminth transcriptomes using HelmDB: implications for biology and biotechnology.Biotechnol Adv. 2013 Dec;31(8):1109-19. doi: 10.1016/j.biotechadv.2012.12.004. Epub 2012 Dec 21. Biotechnol Adv. 2013. PMID: 23266393 Review.
-
Tetrahymena as a Unicellular Model Eukaryote: Genetic and Genomic Tools.Genetics. 2016 Jun;203(2):649-65. doi: 10.1534/genetics.114.169748. Genetics. 2016. PMID: 27270699 Free PMC article. Review.
Cited by
-
Inferring gene-pathway associations from consolidated transcriptome datasets: an interactive gene network explorer for Tetrahymena thermophila.bioRxiv [Preprint]. 2024 Dec 17:2024.12.12.627356. doi: 10.1101/2024.12.12.627356. bioRxiv. 2024. Update in: NAR Genom Bioinform. 2025 May 27;7(2):lqaf067. doi: 10.1093/nargab/lqaf067. PMID: 39713406 Free PMC article. Updated. Preprint.
-
Proteins that control the geometry of microtubules at the ends of cilia.J Cell Biol. 2018 Dec 3;217(12):4298-4313. doi: 10.1083/jcb.201804141. Epub 2018 Sep 14. J Cell Biol. 2018. PMID: 30217954 Free PMC article.
-
An apical membrane complex for triggering rhoptry exocytosis and invasion in Toxoplasma.EMBO J. 2022 Nov 17;41(22):e111158. doi: 10.15252/embj.2022111158. Epub 2022 Oct 17. EMBO J. 2022. PMID: 36245278 Free PMC article.
-
Inferring gene-pathway associations from consolidated transcriptome datasets: an interactive gene network explorer for Tetrahymena thermophila.NAR Genom Bioinform. 2025 May 27;7(2):lqaf067. doi: 10.1093/nargab/lqaf067. eCollection 2025 Jun. NAR Genom Bioinform. 2025. PMID: 40432793 Free PMC article.
References
-
- Witzany G, Nowacki M, editors. Biocommunication of Ciliates. Springer; 2016.
-
- Greider CW, Blackburn EH. Identification of a specific telomere terminal transferase activity in tetrahymena extracts. Cell. 1985;43(2):405– 413. doi: http://dx.doi.org/10.1016/0092-8674(85)90170-9. - DOI - PubMed
-
- Gibbons I, Rowe A. Dynein: a protein with adenosine triphosphatase activity from cilia. Science. 1965;149(3682):424–426. - PubMed
-
- Brownell JE, Zhou J, Ranalli T, Kobayashi R, Edmondson DG, Roth SY, Allis C. Tetrahymena histone acetyltransferase a: A homolog to yeast gcn5p linking histone acetylation to gene activation. Cell. 1996;84(6):843– 851. doi: http://dx.doi.org/10.1016/S0092-8674(00)81063-6. - DOI - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
