An operational view of intercellular signaling pathways
- PMID: 29104946
- PMCID: PMC5665397
- DOI: 10.1016/j.coisb.2016.12.003
An operational view of intercellular signaling pathways
Abstract
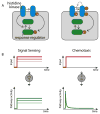
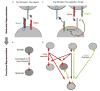
Animal cells use a conserved repertoire of intercellular signaling pathways to communicate with one another. These pathways are well-studied from a molecular point of view. However, we often lack an "operational" understanding that would allow us to use these pathways to rationally control cellular behaviors. This requires knowing what dynamic input features each pathway perceives and how it processes those inputs to control downstream processes. To address these questions, researchers have begun to reconstitute signaling pathways in living cells, analyzing their dynamic responses to stimuli, and developing new functional representations of their behavior. Here we review important insights obtained through these new approaches, and discuss challenges and opportunities in understanding signaling pathways from an operational point of view.
Figures


References
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
