Correlative mass spectrometry imaging, applying time-of-flight secondary ion mass spectrometry and atmospheric pressure matrix-assisted laser desorption/ionization to a single tissue section
- PMID: 29105220
- PMCID: PMC5838509
- DOI: 10.1002/rcm.8022
Correlative mass spectrometry imaging, applying time-of-flight secondary ion mass spectrometry and atmospheric pressure matrix-assisted laser desorption/ionization to a single tissue section
Abstract
Rationale: Mass spectrometry imaging (MSI) is a powerful tool for mapping the surface of a sample. Time-of-flight secondary ion mass spectrometry (TOF-SIMS) and atmospheric pressure matrix-assisted laser desorption/ionization (AP-MALDI) offer complementary capabilities. Here, we present a workflow to apply both techniques to a single tissue section and combine the resulting data for the example of human colon cancer tissue.
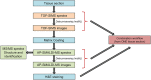
Methods: Following cryo-sectioning, images were acquired using the high spatial resolution (1 μm pixel size) provided by TOF-SIMS. The same section was then coated with a para-nitroaniline matrix and images were acquired using AP-MALDI coupled to an Orbitrap mass spectrometer, offering high mass resolution, high mass accuracy and tandem mass spectrometry (MS/MS) capabilities. Datasets provided by both mass spectrometers were converted into the open and vendor-independent imzML file format and processed with the open-source software MSiReader.
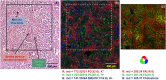
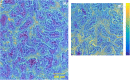
Results: The TOF-SIMS and AP-MALDI mass spectra show strong signals of fatty acids, cholesterol, phosphatidylcholine and sphingomyelin. We showed a high correlation between the fatty acid ions detected with TOF-SIMS in negative ion mode and the phosphatidylcholine ions detected with AP-MALDI in positive ion mode using a similar setting for visualization. Histological staining on the same section allowed the identification of the anatomical structures and their correlation with the ion images.
Conclusions: This multimodal approach using two MSI platforms shows an excellent complementarity for the localization and identification of lipids. The spatial resolution of both systems is at or close to cellular dimensions, and thus spatial correlation can only be obtained if the same tissue section is analyzed sequentially. Data processing based on imzML allows a real correlation of the imaging datasets provided by these two technologies and opens the way for a more complete molecular view of the anatomical structures of biological tissues.
© 2017 The Authors. Rapid Communications in Mass Spectrometry Published by John Wiley & Sons Ltd.
Figures

References
-
- Pól J, Strohalm M, Havlíček V, Volnỷ M. Molecular mass spectrometry imaging in biomedical and life science research. Histochem Cell Biol. 2010;134(5):423‐443. - PubMed
-
- Weaver EM, Hummon AB. Imaging mass spectrometry: From tissue sections to cell cultures. Adv Drug Deliv Rev. 2013;65(8):1039‐1055. - PubMed
-
- Jungmann JH, Heeren RMA. Emerging technologies in mass spectrometry imaging. J Proteomics. 2012;75(16):5077‐5092. - PubMed
-
- Touboul D, Laprévote O, Brunelle A. Micrometric molecular histology of lipids by mass spectrometry imaging. Curr Opin Chem Biol. 2011;15(5):725‐732. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
