Learning a variational network for reconstruction of accelerated MRI data
- PMID: 29115689
- PMCID: PMC5902683
- DOI: 10.1002/mrm.26977
Learning a variational network for reconstruction of accelerated MRI data
Abstract
Purpose: To allow fast and high-quality reconstruction of clinical accelerated multi-coil MR data by learning a variational network that combines the mathematical structure of variational models with deep learning.
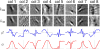
Theory and methods: Generalized compressed sensing reconstruction formulated as a variational model is embedded in an unrolled gradient descent scheme. All parameters of this formulation, including the prior model defined by filter kernels and activation functions as well as the data term weights, are learned during an offline training procedure. The learned model can then be applied online to previously unseen data.
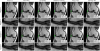
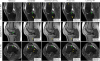
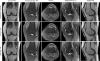
Results: The variational network approach is evaluated on a clinical knee imaging protocol for different acceleration factors and sampling patterns using retrospectively and prospectively undersampled data. The variational network reconstructions outperform standard reconstruction algorithms, verified by quantitative error measures and a clinical reader study for regular sampling and acceleration factor 4.
Conclusion: Variational network reconstructions preserve the natural appearance of MR images as well as pathologies that were not included in the training data set. Due to its high computational performance, that is, reconstruction time of 193 ms on a single graphics card, and the omission of parameter tuning once the network is trained, this new approach to image reconstruction can easily be integrated into clinical workflow. Magn Reson Med 79:3055-3071, 2018. © 2017 International Society for Magnetic Resonance in Medicine.
Keywords: accelerated MRI; compressed sensing; deep learning; image reconstruction; parallel imaging; variational network.
© 2017 International Society for Magnetic Resonance in Medicine.
Figures





References
-
- LeCun Y, Bengio Y, Hinton G. Deep Learning. Nature. 2015;521(7553):436–444. - PubMed
-
- Goodfellow I, Bengio Y, Courville A. Deep Learning. MIT Press; 2016.
-
- Krizhevsky A, Sutskever I, Geoffrey EH. ImageNet Classification with Deep Convolutional Neural Networks. Advances in Neural Information Processing Systems (NIPS) 2012:1097–1105.
-
- Chen LC, Papandreou G, Kokkinos I, Murphy K, Yuille AL. Semantic Image Segmentation with Deep Convolutional Nets and Fully Connected CRFs. International Conference on Learning Representations. 2015:1–14.
-
- Dosovitskiy A, Fischer P, Ilg E, Häusser P, Hazirbas C, Golkov V, van der Smagt P, Cremers D, Brox T. FlowNet: Learning Optical Flow with Convolutional Networks. Proceedings of the IEEE International Conference on Computer Vision (ICCV) 2015:2758–2766.
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

