Small RNAome sequencing delineates the small RNA landscape of pluripotent adult stem cells in the planarian Schmidtea mediterranea
- PMID: 29124009
- PMCID: PMC5671611
- DOI: 10.1016/j.gdata.2017.10.004
Small RNAome sequencing delineates the small RNA landscape of pluripotent adult stem cells in the planarian Schmidtea mediterranea
Abstract
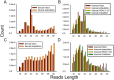
Small noncoding RNAs play a pivotal role in the regulation of gene expression, and are key regulators of animal development. Freshwater planarian exhibits an extraordinary ability to regenerate any missing body parts, representing an emerging model for studying mechanism underlying stem cell regulation and tissue regeneration. Here, we utilized next-generation sequencing (NGS) to identify small RNAs that are expressed in planarian adult stem cells, and are implicated in tissue regeneration. We profiled microRNAs (miRNAs), piwi-interacting RNA (piRNAs), small rDNA-derived RNAs (srRNAs) and endogenous interfering RNAs (endo-siRNAs) population from size 18-30 nt, measured the expression of 244 conserved miRNAs, and identified 41 novel miRNAs and 64 novel endo-siRNAs. Expression profiling analyses revealed that most piRNAs and srRNAs are up-regulated during regeneration, and that the most abundantly expressed srRNAs are from 5.8s and 28s rRNA. Furthermore, a target prediction method was adopted to investigate the anti-correlation of small RNAs and mRNA expression. We built up a gene regulatory network based on the genes that are targeted by dynamically changed small RNAs. These results expand the known small RNA repertoire in planarian, and provide valuable insights and a rich resource for understanding the small RNAs landscape in stem cell-mediated regeneration.
Figures







Similar articles
-
High-resolution profiling and discovery of planarian small RNAs.Proc Natl Acad Sci U S A. 2009 Jul 14;106(28):11546-51. doi: 10.1073/pnas.0905222106. Epub 2009 Jun 29. Proc Natl Acad Sci U S A. 2009. PMID: 19564616 Free PMC article.
-
Dynamic regulation of small RNAome during the early stage of cardiac differentiation from pluripotent embryonic stem cells.Genom Data. 2017 May 3;12:136-145. doi: 10.1016/j.gdata.2017.05.006. eCollection 2017 Jun. Genom Data. 2017. PMID: 28540181 Free PMC article.
-
Identification of small non-coding RNAs in the planarian Dugesia japonica via deep sequencing.Genomics. 2012 May;99(5):315-21. doi: 10.1016/j.ygeno.2012.03.001. Epub 2012 Mar 8. Genomics. 2012. PMID: 22425900
-
Small RNAs in germline development.Curr Top Dev Biol. 2013;102:159-205. doi: 10.1016/B978-0-12-416024-8.00006-4. Curr Top Dev Biol. 2013. PMID: 23287033 Review.
-
Small noncoding RNAs and male infertility.Wiley Interdiscip Rev RNA. 2014 Nov-Dec;5(6):733-45. doi: 10.1002/wrna.1252. Epub 2014 Jul 8. Wiley Interdiscip Rev RNA. 2014. PMID: 25044449 Review.
Cited by
-
Spatiotemporal transcriptomic atlas reveals the dynamic characteristics and key regulators of planarian regeneration.Nat Commun. 2023 Jun 2;14(1):3205. doi: 10.1038/s41467-023-39016-0. Nat Commun. 2023. PMID: 37268637 Free PMC article.
-
Conservation and Targets of miR-71: A Systematic Review and Meta-Analysis.Noncoding RNA. 2023 Jul 26;9(4):41. doi: 10.3390/ncrna9040041. Noncoding RNA. 2023. PMID: 37624033 Free PMC article. Review.
-
miR-8b is involved in brain and eye regeneration of Dugesia japonica in head regeneration.Biol Open. 2021 Jun 15;10(6):bio058538. doi: 10.1242/bio.058538. Epub 2021 Jun 29. Biol Open. 2021. PMID: 34184734 Free PMC article.
-
Efficient Production of On-Target Reads for Small RNA Sequencing of Single Cells Using Modified Adapters.Anal Chem. 2018 Nov 6;90(21):12609-12615. doi: 10.1021/acs.analchem.8b02773. Epub 2018 Sep 27. Anal Chem. 2018. PMID: 30260208 Free PMC article.
-
Comparative Proteome Analysis Indicates The Divergence between The Head and Tail Regeneration in Planarian.Cell J. 2021 Nov;23(6):640-649. doi: 10.22074/cellj.2021.7689. Epub 2021 Nov 23. Cell J. 2021. PMID: 34939757 Free PMC article.
References
-
- Reddien P.W., Alvarado A.S. Fundamentals of planarian regeneration. Annu. Rev. Cell Dev. Biol. 2004;20:725–757. - PubMed
-
- Sánchez Alvarado A. Stem cells in animal models of regeneration. StemBook. 2008:1–24. - PubMed
-
- Liu G., Ding M., Wang H., Huang J., Jing Q., Shen B. Pathway analysis of microRNAs in mouse heart development. Int. J. Bioinforma. Res. Appl. 2010;6:12–20. http://www.ncbi.nlm.nih.gov/pubmed/20110206 - PubMed
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

