ReMap 2018: an updated atlas of regulatory regions from an integrative analysis of DNA-binding ChIP-seq experiments
- PMID: 29126285
- PMCID: PMC5753247
- DOI: 10.1093/nar/gkx1092
ReMap 2018: an updated atlas of regulatory regions from an integrative analysis of DNA-binding ChIP-seq experiments
Abstract
With this latest release of ReMap (http://remap.cisreg.eu), we present a unique collection of regulatory regions in human, as a result of a large-scale integrative analysis of ChIP-seq experiments for hundreds of transcriptional regulators (TRs) such as transcription factors, transcriptional co-activators and chromatin regulators. In 2015, we introduced the ReMap database to capture the genome regulatory space by integrating public ChIP-seq datasets, covering 237 TRs across 13 million (M) peaks. In this release, we have extended this catalog to constitute a unique collection of regulatory regions. Specifically, we have collected, analyzed and retained after quality control a total of 2829 ChIP-seq datasets available from public sources, covering a total of 485 TRs with a catalog of 80M peaks. Additionally, the updated database includes new search features for TR names as well as aliases, including cell line names and the ability to navigate the data directly within genome browsers via public track hubs. Finally, full access to this catalog is available online together with a TR binding enrichment analysis tool. ReMap 2018 provides a significant update of the ReMap database, providing an in depth view of the complexity of the regulatory landscape in human.
© The Author(s) 2017. Published by Oxford University Press on behalf of Nucleic Acids Research.
Figures


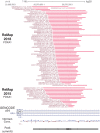
References
-
- Mardis E.R. ChIP-seq: welcome to the new frontier. Nat. Methods. 2007; 4:613–614. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

