NF-κB Dynamics Discriminate between TNF Doses in Single Cells
- PMID: 29128333
- PMCID: PMC5746429
- DOI: 10.1016/j.cels.2017.10.011
NF-κB Dynamics Discriminate between TNF Doses in Single Cells
Abstract
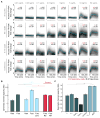
Although cytokine-dependent dynamics of nuclear factor κB (NF-κB) are known to encode information that regulates cell fate decisions, it is unclear whether single-cell responses are switch-like or encode more information about cytokine dose. Here, we measure the dynamic subcellular localization of NF-κB in response to a range of tumor necrosis factor (TNF) stimulation conditions to determine the prevailing mechanism of single-cell dose discrimination. Using an information theory formalism that accounts for signaling dynamics and non-responsive cell subpopulations, we find that the information transmission capacity of single cells exceeds that predicted from a switch-like response. Instead, we observe that NF-κB dynamics within single cells contain sufficient information to encode multiple, TNF-dependent cellular states, and have an activation threshold that varies across the population. By comparing single-cell responses to an internal, experimentally observed reference, we demonstrate that cells can grade responses to TNF across several orders of magnitude in concentration. This suggests that cells contain additional control points to fine-tune their cytokine responses beyond the decision to activate.
Keywords: NF-κB; TNF; information theory; live-cell imaging; signal transduction; single cell; transcription factor.
Copyright © 2017 The Authors. Published by Elsevier Inc. All rights reserved.
Figures
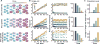


References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

