Comparative transcriptome analysis of soybean response to bean pyralid larvae
- PMID: 29132375
- PMCID: PMC5683215
- DOI: 10.1186/s12864-017-4256-7
Comparative transcriptome analysis of soybean response to bean pyralid larvae
Abstract
Background: Soybean is one of most important oilseed crop worldwide, however, its production is often limited by many insect pests. Bean pyralid is one of the major soybean leaf-feeding insects in China. To explore the defense mechanisms of soybean resistance to bean pyralid, the comparative transcriptome sequencing was completed between the leaves infested with bean pyralid larvae and no worm of soybean (Gantai-2-2 and Wan82-178) on the Illumina HiSeq™ 2000 platform.
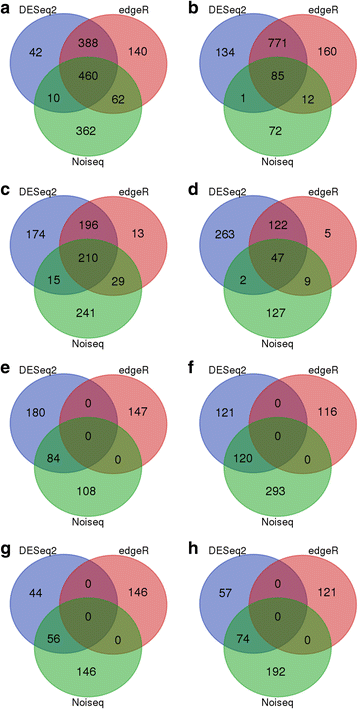
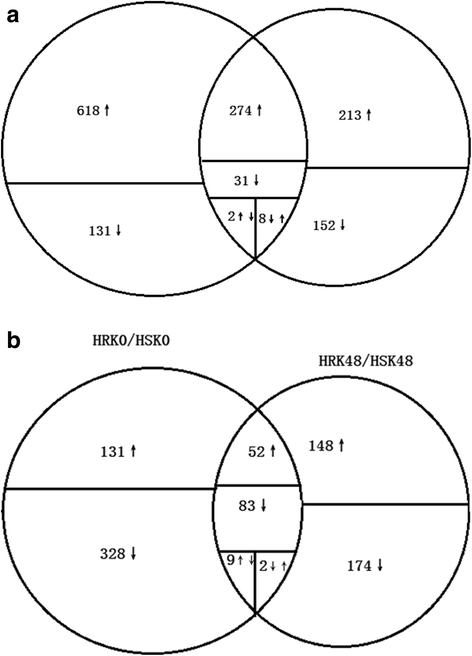
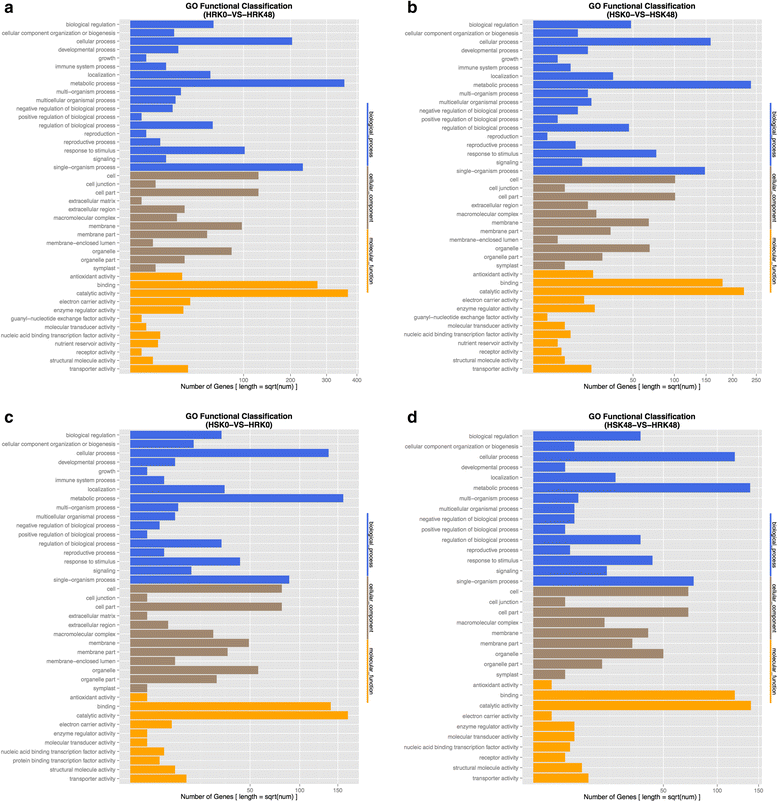
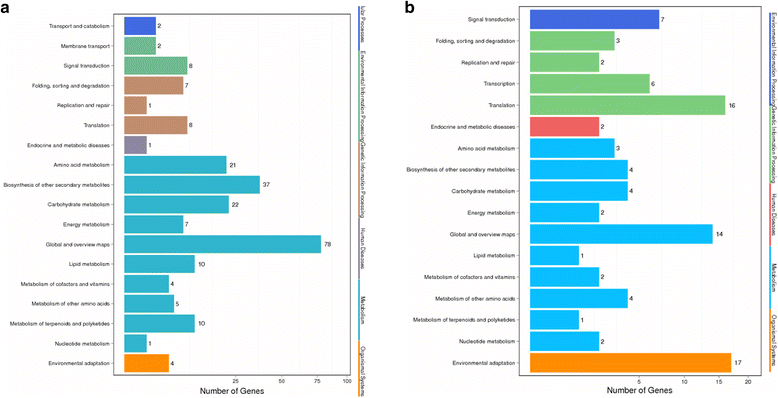
Results: In total, we identified 1744 differentially expressed genes (DEGs) in the leaves of Gantai-2-2 (1064) and Wan82-178 (680) fed by bean pyralid for 48 h, compared to 0 h. Interestingly, 315 DEGs were shared by Gantai-2-2 and Wan82-178, while 749 and 365 DEGs specifically identified in Gantai-2-2 and Wan82-178, respectively. When comparing Gantai-2-2 with Wan82-178, 605 DEGs were identified at 0 h feeding, and 468 DEGs were identified at 48 h feeding. Gene Ontology (GO) annotation analysis revealed that the DEGs were mainly involved in the metabolic process, single-organism process, cellular process, responses to stimulus, catalytic activities and binding. Pathway analysis showed that most of the DEGs were associated with the plant-pathogen interaction, phenylpropanoid biosynthesis, phenylalanine metabolism, flavonoid biosynthesis, peroxisome, plant hormone signal transduction, terpenoid backbone biosynthesis, and so on. Finally, we used qRT-PCR to validate the expression patterns of several genes and the results showed an excellent agreement with deep sequencing.
Conclusions: According to the comparative transcriptome analysis results and related literature reports, we concluded that the response to bean pyralid feeding might be related to the disturbed functions and metabolism pathways of some key DEGs, such as DEGs involved in the ROS removal system, plant hormone metabolism, intracellular signal transduction pathways, secondary metabolism, transcription factors, biotic and abiotic stresses. We speculated that these genes may have played an important role in synthesizing substances to resist insect attacks in soybean. Our results provide a valuable resource of soybean defense genes that will benefit other studies in this field.
Keywords: Bean pyralid; Differentially expressed genes (DEGs); Soybean; Transcriptome sequencing.
Conflict of interest statement
Ethics approval and consent to participate
Not applicable for this research. Soybean seeds for this study were obtained from the Guangxi Academy of Agricultural Science.
Consent for publication
Not applicable
Competing interests
The authors declare that they have no competing interests.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Figures




References
-
- Wlson RF. Chapter 1 soybean; market driven research needs. In: Stacey G, editor. Genetics and genomics of soybean. New York: Springer; 2008. pp. 3–15.
-
- Editorial committee of plate of Chinese diseases and insects on crop . Plate of Chinese diseases and insects on crop, fifth fascicule, diseases and insects on oil crop (first) Beijing: Agricultural press; 1982. pp. 136–137.
-
- Cui ZL, Gai JY, Ji DF, Ren ZJ. A study on leaf-feeding insect species on soybeans in Nanjing area. Soybean Sci. 1997;16(1):12–20.
-
- Sun ZD, Yang SZ, Chen HZ, Li CY, Long LP. Identification of soybean resistance to bean pyralid (Lamprosema indicate Fabricicus) and oviposition preference of bean pyralid on soybean varieties. Chin J Oil Crop Sci. 2005;27(4):69–71.
-
- Sun ZD, Chen HZ, Wei DW. A study on leaf-feeding insect species on soybeans in Nanning. Guangxi Agric Sci. 2001;2:104–106.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

