Spatial Control of Gene Expression by miR319-Regulated TCP Transcription Factors in Leaf Development
- PMID: 29133375
- PMCID: PMC5813565
- DOI: 10.1104/pp.17.00823
Spatial Control of Gene Expression by miR319-Regulated TCP Transcription Factors in Leaf Development
Abstract
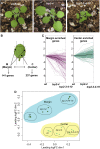
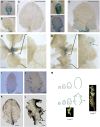
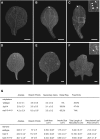
The characteristic leaf shapes we see in all plants are in good part the outcome of the combined action of several transcription factor networks that translate into cell division activity during the early development of the organ. We show here that wild-type leaves have distinct transcriptomic profiles in center and marginal regions. Certain transcripts are enriched in margins, including those of CINCINNATA-like TCPs (TEOSINTE BRANCHED, CYCLOIDEA and PCF1/2) and members of the NGATHA and STYLISH gene families. We study in detail the contribution of microRNA319 (miR319)-regulated TCP transcription factors to the development of the center and marginal regions of Arabidopsis (Arabidopsis thaliana) leaves. We compare in molecular analyses the wild type, the tcp2 tcp4 mutant that has enlarged flat leaves, and the tcp2 tcp3 tcp4 tcp10 mutant with strongly crinkled leaves. The different leaf domains of the tcp mutants show changed expression patterns for many photosynthesis-related genes, indicating delayed differentiation, especially in the marginal parts of the organ. At the same time, we found an up-regulation of cyclin genes and other genes that are known to participate in cell division, specifically in the marginal regions of tcp2 tcp3 tcp4 tcp10 Using GUS reporter constructs, we confirmed extended mitotic activity in the tcp2 tcp3 tcp4 tcp10 leaf, which persisted in small defined foci in the margins when the mitotic activity had already ceased in wild-type leaves. Our results describe the role of miR319-regulated TCP transcription factors in the coordination of activities in different leaf domains during organ development.
© 2018 American Society of Plant Biologists. All Rights Reserved.
Figures



References
-
- Anders S, McCarthy DJ, Chen Y, Okoniewski M, Smyth GK, Huber W, Robinson MD (2013) Count-based differential expression analysis of RNA sequencing data using R and Bioconductor. Nat Protoc 8: 1765–1786 - PubMed
-
- Andriankaja M, Dhondt S, De Bodt S, Vanhaeren H, Coppens F, De Milde L, Mühlenbock P, Skirycz A, Gonzalez N, Beemster GT, et al. (2012) Exit from proliferation during leaf development in Arabidopsis thaliana: a not-so-gradual process. Dev Cell 22: 64–78 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

