Frequent nonallelic gene conversion on the human lineage and its effect on the divergence of gene duplicates
- PMID: 29138319
- PMCID: PMC5715747
- DOI: 10.1073/pnas.1708151114
Frequent nonallelic gene conversion on the human lineage and its effect on the divergence of gene duplicates
Abstract
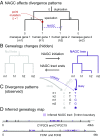
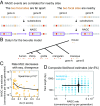
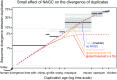
Gene conversion is the copying of a genetic sequence from a "donor" region to an "acceptor." In nonallelic gene conversion (NAGC), the donor and the acceptor are at distinct genetic loci. Despite the role NAGC plays in various genetic diseases and the concerted evolution of gene families, the parameters that govern NAGC are not well characterized. Here, we survey duplicate gene families and identify converted tracts in 46% of them. These conversions reflect a large GC bias of NAGC. We develop a sequence evolution model that leverages substantially more information in duplicate sequences than used by previous methods and use it to estimate the parameters that govern NAGC in humans: a mean converted tract length of 250 bp and a probability of [Formula: see text] per generation for a nucleotide to be converted (an order of magnitude higher than the point mutation rate). Despite this high baseline rate, we show that NAGC slows down as duplicate sequences diverge-until an eventual "escape" of the sequences from its influence. As a result, NAGC has a small average effect on the sequence divergence of duplicates. This work improves our understanding of the NAGC mechanism and the role that it plays in the evolution of gene duplicates.
Keywords: GC bias; gene conversion; gene duplicates; mutation rate; sequence evolution.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

References
-
- Chen JM, Cooper DN, Chuzhanova N, Férec C, Patrinos GP. Gene conversion: Mechanisms, evolution and human disease. Nat Rev Genet. 2007;8:762–775. - PubMed
-
- Innan H, Kondrashov F. The evolution of gene duplications: Classifying and distinguishing between models. Nat Rev Genet. 2010;11:97–108. - PubMed
-
- Bischoff J, et al. Genome-wide identification of pseudogenes capable of disease-causing gene conversion. Hum Mutat. 2006;27:545–552. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

