A comprehensive structural, biochemical and biological profiling of the human NUDIX hydrolase family
- PMID: 29142246
- PMCID: PMC5688067
- DOI: 10.1038/s41467-017-01642-w
A comprehensive structural, biochemical and biological profiling of the human NUDIX hydrolase family
Abstract
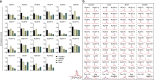
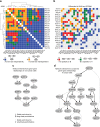
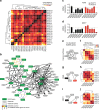
The NUDIX enzymes are involved in cellular metabolism and homeostasis, as well as mRNA processing. Although highly conserved throughout all organisms, their biological roles and biochemical redundancies remain largely unclear. To address this, we globally resolve their individual properties and inter-relationships. We purify 18 of the human NUDIX proteins and screen 52 substrates, providing a substrate redundancy map. Using crystal structures, we generate sequence alignment analyses revealing four major structural classes. To a certain extent, their substrate preference redundancies correlate with structural classes, thus linking structure and activity relationships. To elucidate interdependence among the NUDIX hydrolases, we pairwise deplete them generating an epistatic interaction map, evaluate cell cycle perturbations upon knockdown in normal and cancer cells, and analyse their protein and mRNA expression in normal and cancer tissues. Using a novel FUSION algorithm, we integrate all data creating a comprehensive NUDIX enzyme profile map, which will prove fundamental to understanding their biological functionality.
Conflict of interest statement
The authors declare no competing financial interests.
Figures





Similar articles
-
YZGD from Paenibacillus thiaminolyticus, a pyridoxal phosphatase of the HAD (haloacid dehalogenase) superfamily and a versatile member of the Nudix (nucleoside diphosphate x) hydrolase superfamily.Biochem J. 2006 Mar 15;394(Pt 3):665-74. doi: 10.1042/BJ20051172. Biochem J. 2006. PMID: 16336194 Free PMC article.
-
The evolution of function within the Nudix homology clan.Proteins. 2017 May;85(5):775-811. doi: 10.1002/prot.25223. Epub 2017 Mar 16. Proteins. 2017. PMID: 27936487 Free PMC article.
-
Substrate specificity characterization for eight putative nudix hydrolases. Evaluation of criteria for substrate identification within the Nudix family.Proteins. 2016 Dec;84(12):1810-1822. doi: 10.1002/prot.25163. Epub 2016 Oct 1. Proteins. 2016. PMID: 27618147 Free PMC article.
-
Versatile physiological functions of the Nudix hydrolase family in Arabidopsis.Biosci Biotechnol Biochem. 2015;79(3):354-66. doi: 10.1080/09168451.2014.987207. Epub 2014 Dec 6. Biosci Biotechnol Biochem. 2015. PMID: 25483172 Review.
-
The plant Nudix hydrolase family.Acta Biochim Pol. 2008;55(4):663-71. Epub 2008 Dec 16. Acta Biochim Pol. 2008. PMID: 19081844 Review.
Cited by
-
Fluorescent probe displacement assays reveal unique nucleic acid binding properties of human nudix enzymes.Anal Biochem. 2020 Apr 15;595:113622. doi: 10.1016/j.ab.2020.113622. Epub 2020 Feb 12. Anal Biochem. 2020. PMID: 32059949 Free PMC article.
-
Inositol pyrophosphate catabolism by three families of phosphatases regulates plant growth and development.PLoS Genet. 2024 Nov 12;20(11):e1011468. doi: 10.1371/journal.pgen.1011468. eCollection 2024 Nov. PLoS Genet. 2024. PMID: 39531477 Free PMC article.
-
Uridine-sensitized screening identifies genes and metabolic regulators of nucleotide synthesis.bioRxiv [Preprint]. 2025 Mar 13:2025.03.11.642569. doi: 10.1101/2025.03.11.642569. bioRxiv. 2025. PMID: 40161720 Free PMC article. Preprint.
-
Mitotic MTH1 Inhibitors in Treatment of Cancer.Cancer Treat Res. 2023;186:223-237. doi: 10.1007/978-3-031-30065-3_13. Cancer Treat Res. 2023. PMID: 37978139
-
Host 5'-3' Exoribonuclease XRN1 Acts as a Proviral Factor for Measles Virus Replication by Downregulating the dsRNA-Activated Kinase PKR.J Virol. 2022 Nov 23;96(22):e0131922. doi: 10.1128/jvi.01319-22. Epub 2022 Oct 27. J Virol. 2022. PMID: 36300942 Free PMC article.
References
-
- McLennan AG. The MutT motif family of nucleotide phosphohydrolases in man and human pathogens (review) Int. J. Mol. Med. 1999;4:79–89. - PubMed
-
- Sakumi K, et al. Cloning and expression of cDNA for a human enzyme that hydrolyzes 8-oxo-dGTP, a mutagenic substrate for DNA synthesis. J. Biol. Chem. 1993;268:23524–23530. - PubMed

