Salt-responsive gut commensal modulates TH17 axis and disease
- PMID: 29143823
- PMCID: PMC6070150
- DOI: 10.1038/nature24628
Salt-responsive gut commensal modulates TH17 axis and disease
Abstract
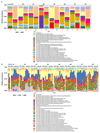
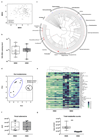
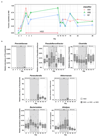
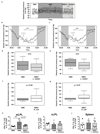
A Western lifestyle with high salt consumption can lead to hypertension and cardiovascular disease. High salt may additionally drive autoimmunity by inducing T helper 17 (TH17) cells, which can also contribute to hypertension. Induction of TH17 cells depends on gut microbiota; however, the effect of salt on the gut microbiome is unknown. Here we show that high salt intake affects the gut microbiome in mice, particularly by depleting Lactobacillus murinus. Consequently, treatment of mice with L. murinus prevented salt-induced aggravation of actively induced experimental autoimmune encephalomyelitis and salt-sensitive hypertension by modulating TH17 cells. In line with these findings, a moderate high-salt challenge in a pilot study in humans reduced intestinal survival of Lactobacillus spp., increased TH17 cells and increased blood pressure. Our results connect high salt intake to the gut-immune axis and highlight the gut microbiome as a potential therapeutic target to counteract salt-sensitive conditions.
Conflict of interest statement
Figures








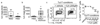

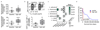
Comment in
-
Microbiota: A high-pressure situation for bacteria.Nature. 2017 Nov 30;551(7682):571-572. doi: 10.1038/nature24760. Epub 2017 Nov 15. Nature. 2017. PMID: 29143820 No abstract available.
-
Hypertension: Salt: the microbiome, immune function and hypertension.Nat Rev Nephrol. 2018 Feb;14(2):71. doi: 10.1038/nrneph.2017.166. Epub 2017 Nov 27. Nat Rev Nephrol. 2018. PMID: 29176658 No abstract available.
References
-
- Taylor J. 2013 ESH/ESC guidelines for the management of arterial hypertension. European heart journal. 2013;34:2108–2109. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Miscellaneous

