Comparison of ChIP-Seq Data and a Reference Motif Set for Human KRAB C2H2 Zinc Finger Proteins
- PMID: 29146583
- PMCID: PMC5765350
- DOI: 10.1534/g3.117.300296
Comparison of ChIP-Seq Data and a Reference Motif Set for Human KRAB C2H2 Zinc Finger Proteins
Abstract
KRAB C2H2 zinc finger proteins (KZNFs) are the largest and most diverse family of human transcription factors, likely due to diversifying selection driven by novel endogenous retroelements (EREs), but the vast majority lack binding motifs or functional data. Two recent studies analyzed a majority of the human KZNFs using either ChIP-seq (60 proteins) or ChIP-exo (221 proteins) in the same cell type (HEK293). The ChIP-exo paper did not describe binding motifs, however. Thirty-nine proteins are represented in both studies, enabling the systematic comparison of the data sets presented here. Typically, only a minority of peaks overlap, but the two studies nonetheless display significant similarity in ERE binding for 32/39, and yield highly similar DNA binding motifs for 23 and related motifs for 34 (MoSBAT similarity score >0.5 and >0.2, respectively). Thus, there is overall (albeit imperfect) agreement between the two studies. For the 242 proteins represented in at least one study, we selected a highest-confidence motif for each protein, utilizing several motif-derivation approaches, and evaluating motifs within and across data sets. Peaks for the majority (158) are enriched (96% with AUC >0.6 predicting peak vs. nonpeak) for a motif that is supported by the C2H2 "recognition code," consistent with intrinsic sequence specificity driving DNA binding in cells. An additional 63 yield motifs enriched in peaks, but not supported by the recognition code, which could reflect indirect binding. Altogether, these analyses validate both data sets, and provide a reference motif set with associated quality metrics.
Keywords: C2H2 recognition code; ChIP-seq; DNA-binding motif; KRAB C2H2 zinc finger proteins; endogenous retroelements.
Copyright © 2018 Barazandeh et al.
Figures

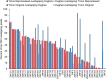





References
-
- Brayer K. J., Segal D. J., 2008. Keep your fingers off my DNA: protein-protein interactions mediated by C2H2 zinc finger domains. Cell Biochem Biophys 50: 111–131. - PubMed
-
- Deplancke B., Alpern D., Gardeux V., 2016. The genetics of transcription factor DNA binding variation. Cell 166: 538–554. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
