Defense against Reactive Carbonyl Species Involves at Least Three Subcellular Compartments Where Individual Components of the System Respond to Cellular Sugar Status
- PMID: 29150548
- PMCID: PMC5757266
- DOI: 10.1105/tpc.17.00258
Defense against Reactive Carbonyl Species Involves at Least Three Subcellular Compartments Where Individual Components of the System Respond to Cellular Sugar Status
Abstract
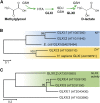
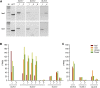
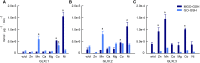
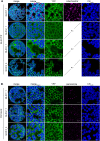
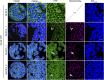
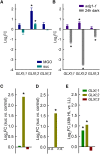
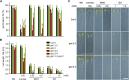
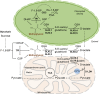
Methylglyoxal (MGO) and glyoxal (GO) are toxic reactive carbonyl species generated as by-products of glycolysis. The pre-emption pathway for detoxification of these products, the glyoxalase (GLX) system, involves two consecutive reactions catalyzed by GLXI and GLXII. In Arabidopsis thaliana, the GLX system is encoded by three homologs of GLXI and three homologs of GLXII, from which several predicted GLXI and GLXII isoforms can be derived through alternative splicing. We identified the physiologically relevant splice forms using sequencing data and demonstrated that the resulting isoforms have different subcellular localizations. All three GLXI homologs are functional in vivo, as they complemented a yeast GLXI loss-of-function mutant. Efficient MGO and GO detoxification can be controlled by a switch in metal cofactor usage. MGO formation is closely connected to the flux through glycolysis and through the Calvin Benson cycle; accordingly, expression analysis indicated that GLXI is transcriptionally regulated by endogenous sugar levels. Analyses of Arabidopsis loss-of-function lines revealed that the elimination of toxic reactive carbonyl species during germination and seedling establishment depends on the activity of the cytosolic GLXI;3 isoform. The Arabidopsis GLX system involves the cytosol, chloroplasts, and mitochondria, which harbor individual components that might be used at specific developmental stages and respond differentially to cellular sugar status.
© 2017 American Society of Plant Biologists. All rights reserved.
Figures


References
-
- Aldini G., Vistoli G., Stefek M., Chondrogianni N., Grune T., Sereikaite J., Sadowska-Bartosz I., Bartosz G. (2013). Molecular strategies to prevent, inhibit, and degrade advanced glycoxidation and advanced lipoxidation end products. Free Radic. Res. 47 (suppl. 1): 93–137. - PubMed
-
- Arrivault S., Guenther M., Ivakov A., Feil R., Vosloh D., van Dongen J.T., Sulpice R., Stitt M. (2009). Use of reverse-phase liquid chromatography, linked to tandem mass spectrometry, to profile the Calvin cycle and other metabolic intermediates in Arabidopsis rosettes at different carbon dioxide concentrations. Plant J. 59: 826–839. - PubMed
-
- Bechtold U., Rabbani N., Mullineaux P.M., Thornalley P.J. (2009). Quantitative measurement of specific biomarkers for protein oxidation, nitration and glycation in Arabidopsis leaves. Plant J. 59: 661–671. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

