Pan-Cellulosomics of Mesophilic Clostridia: Variations on a Theme
- PMID: 29156585
- PMCID: PMC5748583
- DOI: 10.3390/microorganisms5040074
Pan-Cellulosomics of Mesophilic Clostridia: Variations on a Theme
Abstract
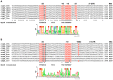
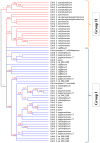
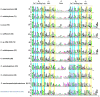
The bacterial cellulosome is an extracellular, multi-enzyme machinery, which efficiently depolymerizes plant biomass by degrading plant cell wall polysaccharides. Several cellulolytic bacteria have evolved various elaborate modular architectures of active cellulosomes. We present here a genome-wide analysis of a dozen mesophilic clostridia species, including both well-studied and yet-undescribed cellulosome-producing bacteria. We first report here, the presence of cellulosomal elements, thus expanding our knowledge regarding the prevalence of the cellulosomal paradigm in nature. We explored the genomic organization of key cellulosome components by comparing the cellulosomal gene clusters in each bacterial species, and the conserved sequence features of the specific cellulosomal modules (cohesins and dockerins), on the background of their phylogenetic relationship. Additionally, we performed comparative analyses of the species-specific repertoire of carbohydrate-degrading enzymes for each of the clostridial species, and classified each cellulosomal enzyme into a specific CAZy family, thus indicating their putative enzymatic activity (e.g., cellulases, hemicellulases, and pectinases). Our work provides, for this large group of bacteria, a broad overview of the blueprints of their multi-component cellulosomal complexes. The high similarity of their scaffoldin clusters and dockerin-based recognition residues suggests a common ancestor, and/or extensive horizontal gene transfer, and potential cross-species recognition. In addition, the sporadic spatial organization of the numerous dockerin-containing genes in several of the genomes, suggests the importance of the cellulosome paradigm in the given bacterial species. The information gained in this work may be utilized directly or developed further by genetically engineering and optimizing designer cellulosome systems for enhanced biotechnological biomass deconstruction and biofuel production.
Keywords: cellulosomes; cohesin; dockerin; glycoside hydrolases; scaffoldin.
Conflict of interest statement
The authors declare no conflict of interest.
Figures




Similar articles
-
Elaborate cellulosome architecture of Acetivibrio cellulolyticus revealed by selective screening of cohesin-dockerin interactions.PeerJ. 2014 Oct 30;2:e636. doi: 10.7717/peerj.636. eCollection 2014. PeerJ. 2014. PMID: 25374780 Free PMC article.
-
Unique organization and unprecedented diversity of the Bacteroides (Pseudobacteroides) cellulosolvens cellulosome system.Biotechnol Biofuels. 2017 Sep 7;10:211. doi: 10.1186/s13068-017-0898-6. eCollection 2017. Biotechnol Biofuels. 2017. PMID: 28912832 Free PMC article.
-
Minimalistic Cellulosome of the Butanologenic Bacterium Clostridium saccharoperbutylacetonicum.mBio. 2020 Mar 31;11(2):e00443-20. doi: 10.1128/mBio.00443-20. mBio. 2020. PMID: 32234813 Free PMC article.
-
Cellulosomes: Highly Efficient Cellulolytic Complexes.Subcell Biochem. 2021;96:323-354. doi: 10.1007/978-3-030-58971-4_9. Subcell Biochem. 2021. PMID: 33252735 Review.
-
Cellulosomes - promising supramolecular machines of anaerobic cellulolytic microorganisms.Acta Chim Slov. 2010 Dec;57(4):767-74. Acta Chim Slov. 2010. PMID: 24061876 Review.
Cited by
-
A genomic analysis reveals the diversity of cellulosome displaying bacteria.Front Microbiol. 2024 Oct 30;15:1473396. doi: 10.3389/fmicb.2024.1473396. eCollection 2024. Front Microbiol. 2024. PMID: 39539715 Free PMC article.
-
Phylogenomics and genetic analysis of solvent-producing Clostridium species.Sci Data. 2024 May 1;11(1):432. doi: 10.1038/s41597-024-03210-6. Sci Data. 2024. PMID: 38693191 Free PMC article.
-
Insight into the autoproteolysis mechanism of the RsgI9 anti-σ factor from Clostridium thermocellum.Proteins. 2024 Aug;92(8):946-958. doi: 10.1002/prot.26690. Epub 2024 Apr 10. Proteins. 2024. PMID: 38597224 Free PMC article.
-
Mapping the deformability of natural and designed cellulosomes in solution.Biotechnol Biofuels Bioprod. 2022 Jun 20;15(1):68. doi: 10.1186/s13068-022-02165-3. Biotechnol Biofuels Bioprod. 2022. PMID: 35725490 Free PMC article.
-
A cellulosomal double-dockerin module from Clostridium thermocellum shows distinct structural and cohesin-binding features.Protein Sci. 2024 Apr;33(4):e4937. doi: 10.1002/pro.4937. Protein Sci. 2024. PMID: 38501488 Free PMC article.
References
-
- Albersheim P., Darvill A., Roberts K., Sederoff R., Staehelin A. Plant Cell Walls. Garland Science; New York, NY, USA: 2010. p. 430.
-
- Himmel M.E., Xu Q., Luo Y., Ding S.-Y., Lamed R., Bayer E.A. Microbial enzyme systems for biomass conversion: Emerging paradigms. Biofuels. 2010;1:323–341. doi: 10.4155/bfs.09.25. - DOI
-
- Bayer E.A., Shoham Y., Lamed R. Lignocellulose-decomposing bacteria and their enzyme systems. In: Rosenberg E., editor. The Prokaryotes. 4th ed. Springer; Berlin, Germany: 2013. pp. 216–266.
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

